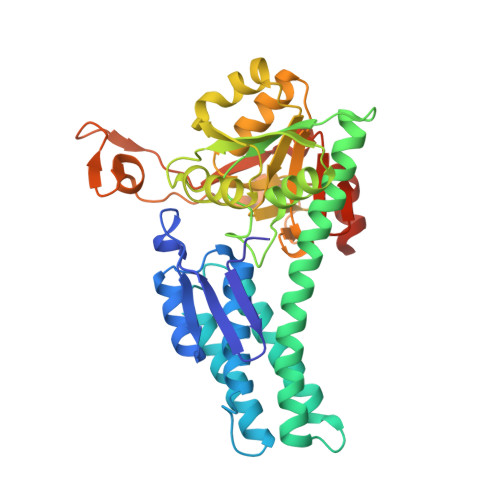
Structure of Mediator of RhoA-Dependent Invasion (MRDI) Explains Its Dual Function as a Metabolic Enzyme and a Mediator of Cell Invasion.
Templeton, P.D., Litman, E.S., Metzner, S.I., Ahn, N.G., Sousa, M.C.(2013) Biochemistry 52: 5675-5684
- PubMed: 23859498
- DOI: https://doi.org/10.1021/bi400556e
- Primary Citation of Related Structures:
4LDQ, 4LDR - PubMed Abstract:
Metastatic melanoma is among the most intractable cancers to treat; patients show resistance to therapy and limited survival time. A critical step in the development of metastatic melanoma is the acquisition of invasion and transition from thin to thick tumors on the skin, followed by invasion to lymph nodes. Prior studies have shown that metastatic melanoma is associated with dysregulation of RhoA and enhanced expression of a protein named "mediator of RhoA-dependent invasion (MRDI)". Importantly, MRDI is a "moonlighting" enzyme, with two distinct functions in melanoma cells. First, MRDI acts as a methylthioribose-1-phosphate (MTR-1-P) isomerase, catalyzing a critical step in methionine salvage. Second, MRDI promotes and is necessary for melanoma cell invasion, independent of its catalytic activity. This paper demonstrates that MtnA, a bacterial MTR-1-P isomerase, rescues the methionine salvage function of MRDI, but is unable to rescue its role in invasion. The crystal structure of MRDI was solved to a resolution of 2.5 Å to identify structural elements important for its invasion activity. This structure and its comparison with other MTR-1-P isomerases are presented, and mutations within a region separate from the MTR-1-P binding site, which interfere with invasion, are identified. Thus, structural elements in MRDI distal from the MTR-1-P catalytic site are responsible for the invasion phenotype.
- Department of Chemistry and Biochemistry, University of Colorado at Boulder, Boulder, CO 80309-0596, USA.
Organizational Affiliation: