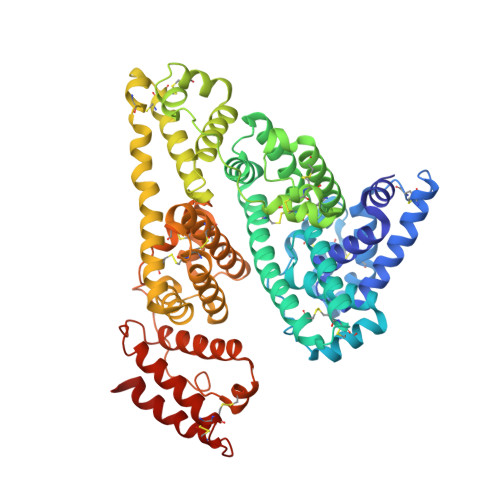
Structural studies of several clinically important oncology drugs in complex with human serum albumin.
Wang, Z.M., Ho, J.X., Ruble, J.R., Rose, J., Ruker, F., Ellenburg, M., Murphy, R., Click, J., Soistman, E., Wilkerson, L., Carter, D.C.(2013) Biochim Biophys Acta 1830: 5356-5374
- PubMed: 23838380 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbagen.2013.06.032
- Primary Citation Related Structures:
4L8U, 4L9K, 4L9Q, 4LA0, 4LB2, 4LB9 - PubMed Abstract:
Serum albumin is a major pharmacokinetic effector of drugs. To gain further insight into albumin binding chemistry, the crystal structures of six oncology agents were determined in complex with human serum albumin at resolutions of 2.8 to 2.0Å: camptothecin, 9-amino-camptothecin, etoposide, teniposide, bicalutamide and idarubicin. Protein crystal growth and low temperature X-ray crystallography These large, complex drugs are all bound within the subdomain IB binding region which can be described as a hydrophobic groove formed by α-helices h7, h8 and h9 covered by the extended polypeptide L1. L1 creates a binding cavity with two access sites, one between loop L1 and α-helices h7 and h8 (distal site: IBd) and the other between L1 and α-helix h9 (proximal site: IBp). Camptothecin (2.4Å) and 9 amino camptothecin (2.0Å) are clearly bound as the open lactone form (IBp). Idarubicin (2.8Å) binds in a DNA like dimer complex via an intermolecular π stacking arrangement in IBd. Bicalutamide (2.4Å) is bound in a folded intramolecular π stacking arrangement between two aromatic rings in IBd similar to idarubicin. Teniposide (2.7Å) and etoposide (2.7Å), despite small chemical differences, are bound in two distinctly different sites at or near IB. Teniposide is internalized via primarily hydrophobic interactions and spans through both openings (IBp-d). Etoposide is bound between the exterior of IB and IIA and exhibits an extensive hydrogen bonding network. Subdomain IB is a major binding site for complex heterocyclic molecules. The structures have important implications for drug design and development. This article is part of a Special Issue entitled Serum Albumin.
- New Century Pharmaceuticals, Inc., 895 Martin Road, Suite C, Huntsville, AL 35824, USA.
Organizational Affiliation: