NMR binding and crystal structure reveal that intrinsically-unstructured regulatory domain auto-inhibits PAK4 by a mechanism different for that of PAK1
Wang, W., Lim, L., Baskaran, Y., Manser, E., Song, J.(2013) Biochem Biophys Res Commun 438: 169-174
- PubMed: 23876315
- DOI: https://doi.org/10.1016/j.bbrc.2013.07.047
- Primary Citation of Related Structures:
4L67 - PubMed Abstract:
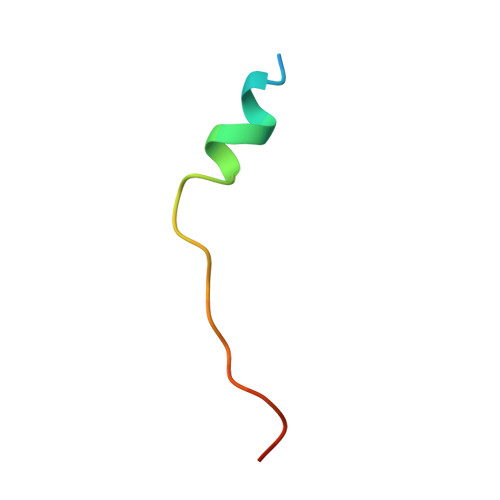
Six human PAK members are classified into groups I (PAKs 1-3) and II (PAK4-6). Previously, only group I PAKs were thought to be auto-inhibited but very recently PAK4, the prototype of group II PAKs, has also been shown to be auto-inhibited by its N-terminal regulatory domain. However, the complete auto-inhibitory domain (AID) sequence remains undefined and the mechanism underlying its auto-inhibition is largely elusive. Here, the N-terminal regulatory domain of PAK4 sufficient for auto-inhibiting and binding Cdc42/Rac was characterized to be intrinsically unstructured, but nevertheless we identified the entire AID sequence by NMR. Strikingly, an AID peptide was derived by deleting the binding-unnecessary residues, which has a Kd of 320 nM to the PAK4 catalytic domain. Consequently, the PAK4 crystal structure complexed with the entire AID has been determined, which reveals that the complete kinase cleft is occupied by 20 AID residuescomposed of an N-terminal α-helix and a previously-identified pseudosubstrate motif, thus achieving auto-inhibition. Our study reveals that PAK4 is auto-inhibited by a novel mechanism which is completely different from that for PAK1, thus bearing critical implications for design of inhibitors specific for group II PAKs.
- Department of Biological Sciences, Faculty of Science, National University of Singapore, 10 Kent Ridge Crescent, Singapore 119260, Singapore.
Organizational Affiliation: