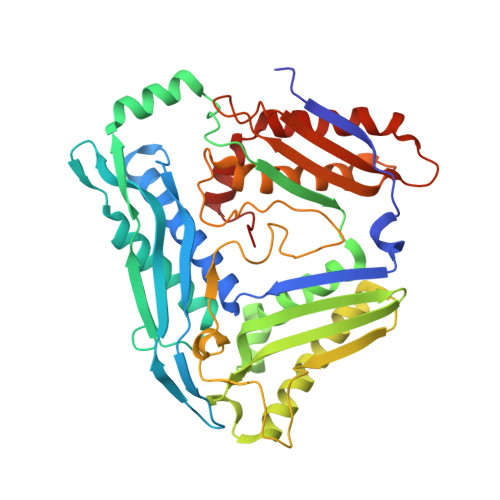
Structural and functional characterisation of the methionine adenosyltransferase from Thermococcus kodakarensis.
Schlesier, J., Siegrist, J., Gerhardt, S., Erb, A., Blaesi, S., Richter, M., Einsle, O., Andexer, J.N.(2013) BMC Struct Biol 13: 22-22
- PubMed: 24134203
- DOI: https://doi.org/10.1186/1472-6807-13-22
- Primary Citation Related Structures:
4L4Q - PubMed Abstract:
Methionine adenosyltransferases catalyse the synthesis of S-adenosylmethionine, a cofactor abundant in all domains of life. In contrast to the enzymes from bacteria and eukarya that show high sequence similarity, methionine adenosyltransferases from archaea diverge on the amino acid sequence level and only few conserved residues are retained. We describe the initial characterisation and the crystal structure of the methionine adenosyltransferase from the hyperthermophilic archaeon Thermococcus kodakarensis. As described for other archaeal methionine adenosyltransferases the enzyme is a dimer in solution and shows high temperature stability. The overall structure is very similar to that of the bacterial and eukaryotic enzymes described, with some additional features that might add to the stability of the enzyme. Compared to bacterial and eukaryotic structures, the active site architecture is largely conserved, with some variation in the substrate/product-binding residues. A flexible loop that was not fully ordered in previous structures without ligands in the active side is clearly visible and forms a helix that leaves an entrance to the active site open. The similar three-dimensional structures of archaeal and bacterial or eukaryotic methionine adenosyltransferases support that these enzymes share an early common ancestor from which they evolved independently, explaining the low similarity in their amino acid sequences. Furthermore, methionine adenosyltransferase from T. kodakarensis is the first structure without any ligands bound in the active site where the flexible loop covering the entrance to the active site is fully ordered, supporting a mechanism postulated earlier for the methionine adenosyltransferase from E. coli. The structure will serve as a starting point for further mechanistic studies and permit the generation of enzyme variants with different characteristics by rational design.
- Institute of Pharmaceutical Sciences, Albert-Ludwigs-University Freiburg, Albertstr, 25, Freiburg D-79104, Germany. jennifer.andexer@pharmazie.uni-freiburg.de.
Organizational Affiliation: