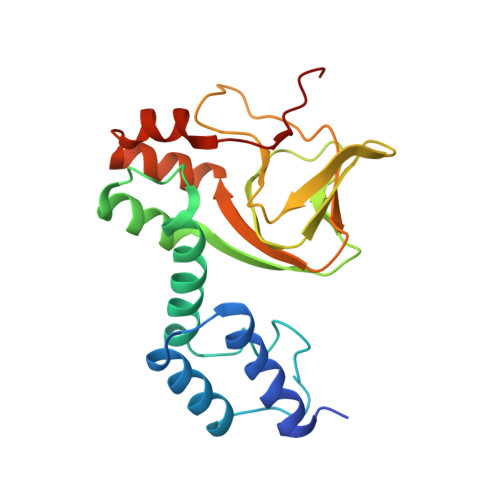
Structure of the C-terminal region of an ERG channel and functional implications.
Brelidze, T.I., Gianulis, E.C., Dimaio, F., Trudeau, M.C., Zagotta, W.N.(2013) Proc Natl Acad Sci U S A 110: 11648-11653
- PubMed: 23801759
- DOI: https://doi.org/10.1073/pnas.1306887110
- Primary Citation Related Structures:
4L11 - PubMed Abstract:
The human ether-à-go-go-related gene (hERG) encodes a K(+) channel crucial for repolarization of the cardiac action potential. EAG-related gene (ERG) channels contain a C-terminal cyclic nucleotide-binding homology domain coupled to the pore of the channel by a C-linker. Here, we report the structure of the C-linker/cyclic nucleotide-binding homology domain of a mosquito ERG channel at 2.5-Å resolution. The structure reveals that the region expected to form the cyclic nucleotide-binding pocket is negatively charged and is occupied by a short β-strand, referred to as the intrinsic ligand, explaining the lack of direct regulation of ERG channels by cyclic nucleotides. In hERG channels, the intrinsic ligand harbors hereditary mutations associated with long-QT syndrome (LQTS), a potentially lethal cardiac arrhythmia. Mutations in the intrinsic ligand affected hERG channel gating and LQTS mutations abolished hERG currents and altered trafficking of hERG channels, which explains the LQT phenotype. The structure also reveals a dramatically different conformation of the C-linker compared with the structures of the related ether-à-go-go-like K(+) and hyperpolarization-activated cyclic nucleotide-modulated channels, suggesting that the C-linker region may be highly dynamic in the KCNH, hyperpolarization-activated cyclic nucleotide-modulated, and cyclic nucleotide-gated channels.
- Department of Physiology and Biophysics, University of Washington School of Medicine, Seattle, WA 98195, USA.
Organizational Affiliation: