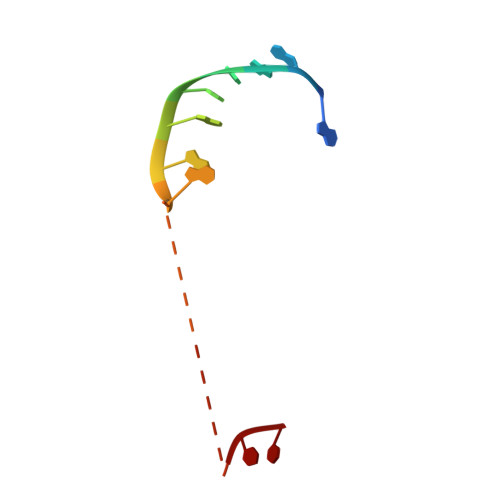
Eukaryote-Specific Insertion Elements Control Human ARGONAUTE Slicer Activity.
Nakanishi, K., Ascano, M., Gogakos, T., Ishibe-Murakami, S., Serganov, A.A., Briskin, D., Morozov, P., Tuschl, T., Patel, D.J.(2013) Cell Rep 3: 1893-1900
- PubMed: 23809764 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2013.06.010
- Primary Citation Related Structures:
4KXT - PubMed Abstract:
We have solved the crystal structure of human ARGONAUTE1 (hAGO1) bound to endogenous 5'-phosphorylated guide RNAs. To identify changes that evolutionarily rendered hAGO1 inactive, we compared our structure with guide-RNA-containing and cleavage-active hAGO2. Aside from mutation of a catalytic tetrad residue, proline residues at positions 670 and 675 in hAGO1 introduce a kink in the cS7 loop, forming a convex surface within the hAGO1 nucleic-acid-binding channel near the inactive catalytic site. We predicted that even upon restoration of the catalytic tetrad, hAGO1-cS7 sterically hinders the placement of a fully paired guide-target RNA duplex into the endonuclease active site. Consistent with this hypothesis, reconstitution of the catalytic tetrad with R805H led to low-level hAGO1 cleavage activity, whereas combining R805H with cS7 substitutions P670S and P675Q substantially augmented hAGO1 activity. Evolutionary amino acid changes to hAGO1 were readily reversible, suggesting that loading of guide RNA and pairing of seed-based miRNA and target RNA constrain its sequence drift.
- Structural Biology Program, Memorial Sloan-Kettering Cancer Center, New York, NY 10065, USA.
Organizational Affiliation: