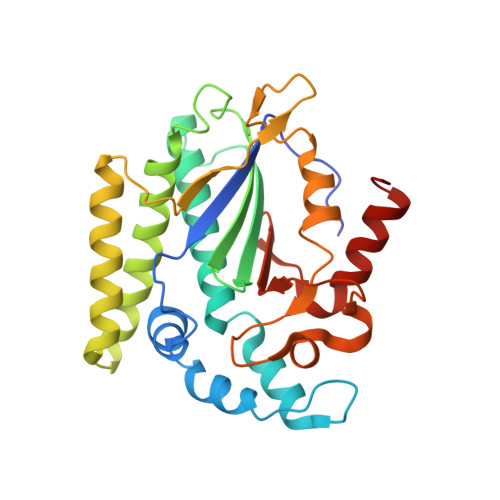
Structure of the NS1 Protein N-Terminal Origin Recognition/Nickase Domain from the Emerging Human Bocavirus.
Tewary, S.K., Zhao, H., Shen, W., Qiu, J., Tang, L.(2013) J Virol 87: 11487-11493
- PubMed: 23966383 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.01770-13
- Primary Citation Related Structures:
4KW3 - PubMed Abstract:
Human bocavirus is a newly identified, globally prevalent, parvovirus that is associated with respiratory infection in infants and young children. Parvoviruses encode a large nonstructural protein 1 (NS1) that is essential for replication of the viral single-stranded DNA genome and DNA packaging and may play versatile roles in virus-host interactions. Here, we report the structure of the human bocavirus NS1 N-terminal domain, the first for any autonomous parvovirus. The structure shows an overall fold that is canonical to the histidine-hydrophobic-histidine superfamily of nucleases, which integrates two distinct DNA-binding sites: (i) a positively charged region mediated by a surface hairpin (residues 190 to 198) that is responsible for recognition of the viral origin of replication of the double-stranded DNA nature and (ii) the nickase active site that binds to the single-stranded DNA substrate for site-specific cleavage. The structure reveals an acidic-residue-rich subdomain that is present in bocavirus NS1 proteins but not in the NS1 orthologs in erythrovirus or dependovirus, which may mediate bocavirus-specific interaction with DNA or potential host factors. These results provide insights into recognition of the origin of replication and nicking of DNA during bocavirus genome replication. Mapping of variable amino acid residues of NS1s from four human bocavirus species onto the structure shows a scattered pattern, but the origin recognition site and the nuclease active site are invariable, suggesting potential targets for antivirals against this clade of highly diverse human viruses.
- Department of Molecular Biosciences, University of Kansas, Lawrence, Kansas, USA.
Organizational Affiliation: