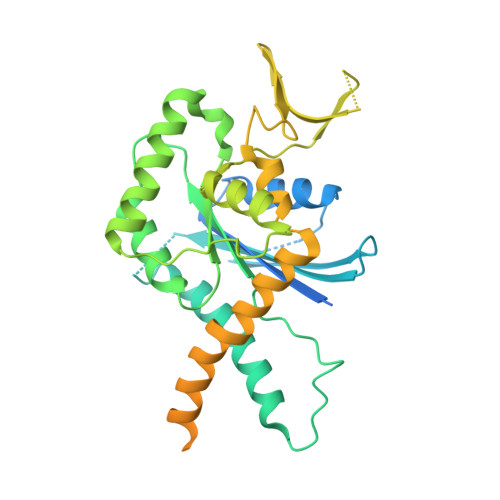
Crystal Structure of a Schistosoma mansoni Septin Reveals the Phenomenon of Strand Slippage in Septins Dependent on the Nature of the Bound Nucleotide.
Zeraik, A.E., Pereira, H.M., Santos, Y.V., Brandao-Neto, J., Spoerner, M., Santos, M.S., Colnago, L.A., Garratt, R.C., Araujo, A.P., Demarco, R.(2014) J Biological Chem 289: 7799-7811
- PubMed: 24464615
- DOI: https://doi.org/10.1074/jbc.M113.525352
- Primary Citation of Related Structures:
4KV9, 4KVA - PubMed Abstract:
Septins are filament-forming GTP-binding proteins involved in important cellular events, such as cytokinesis, barrier formation, and membrane remodeling. Here, we present two crystal structures of the GTPase domain of a Schistosoma mansoni septin (SmSEPT10), one bound to GDP and the other to GTP. The structures have been solved at an unprecedented resolution for septins (1.93 and 2.1 Å, respectively), which has allowed for unambiguous structural assignment of regions previously poorly defined. Consequently, we provide a reliable model for functional interpretation and a solid foundation for future structural studies. Upon comparing the two complexes, we observe for the first time the phenomenon of a strand slippage in septins. Such slippage generates a front-back communication mechanism between the G and NC interfaces. These data provide a novel mechanistic framework for the influence of nucleotide binding to the GTPase domain, opening new possibilities for the study of the dynamics of septin filaments.
- From the Instituto de Física de São Carlos, Universidade de São Paulo, 13563-120 São Carlos, São Paulo, Brazil.
Organizational Affiliation: