The Legionella pneumophila kai operon is implicated in stress response and confers fitness in competitive environments.
Loza-Correa, M., Sahr, T., Rolando, M., Daniels, C., Petit, P., Skarina, T., Gomez Valero, L., Dervins-Ravault, D., Honore, N., Savchenko, A., Buchrieser, C.(2014) Environ Microbiol 16: 359-381
- PubMed: 23957615 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/1462-2920.12223
- Primary Citation Related Structures:
4KUN - PubMed Abstract:
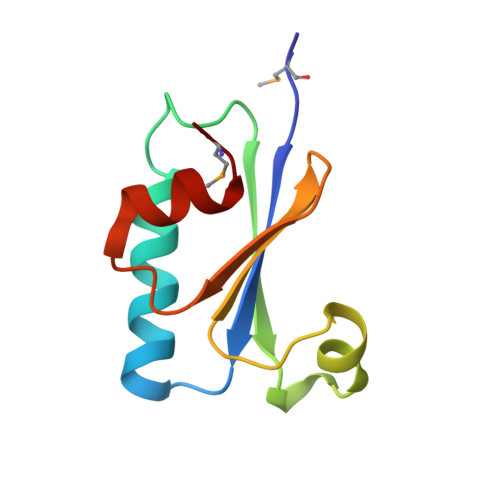
Legionella pneumophila uses aquatic protozoa as replication niche and protection from harsh environments. Although L. pneumophila is not known to have a circadian clock, it encodes homologues of the KaiBC proteins of Cyanobacteria that regulate circadian gene expression. We show that L. pneumophila kaiB, kaiC and the downstream gene lpp1114, are transcribed as a unit under the control of the stress sigma factor RpoS. KaiC and KaiB of L. pneumophila do not interact as evidenced by yeast and bacterial two-hybrid analyses. Fusion of the C-terminal residues of cyanobacterial KaiB to Legionella KaiB restores their interaction. In contrast, KaiC of L. pneumophila conserved autophosphorylation activity, but KaiB does not trigger the dephosphorylation of KaiC like in Cyanobacteria. The crystal structure of L. pneumophila KaiB suggests that it is an oxidoreductase-like protein with a typical thioredoxin fold. Indeed, mutant analyses revealed that the kai operon-encoded proteins increase fitness of L. pneumophila in competitive environments, and confer higher resistance to oxidative and sodium stress. The phylogenetic analysis indicates that L. pneumophila KaiBC resemble Synechosystis KaiC2B2 and not circadian KaiB1C1. Thus, the L. pneumophila Kai proteins do not encode a circadian clock, but enhance stress resistance and adaption to changes in the environments.
- Institut Pasteur, Biologie des Bactéries Intracellulaires, Paris, France; CNRS UMR 3525, Paris, France.
Organizational Affiliation: