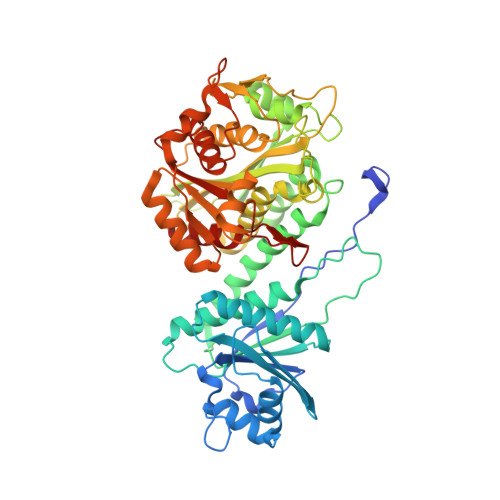
Structure of tomato wound-induced leucine aminopeptidase sheds light on substrate specificity.
Duprez, K., Scranton, M.A., Walling, L.L., Fan, L.(2014) Acta Crystallogr D Biol Crystallogr 70: 1649-1658
- PubMed: 24914976 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004714006245
- Primary Citation Related Structures:
4KSI - PubMed Abstract:
The acidic leucine aminopeptidase (LAP-A) from tomato is induced in response to wounding and insect feeding. Although LAP-A shows in vitro peptidase activity towards peptides and peptide analogs, it is not clear what kind of substrates LAP-A hydrolyzes in vivo. In the current study, the crystal structure of LAP-A was determined to 2.20 Å resolution. Like other LAPs in the M17 peptidase family, LAP-A is a dimer of trimers containing six monomers of bilobal structure. Each monomer contains two metal ions bridged by a water or a hydroxyl ion at the active site. Modeling of different peptides or peptide analogs in the active site of LAP-A reveals a spacious substrate-binding channel that can bind peptides of five or fewer residues with few geometric restrictions. The sequence specificity of the bound peptide is likely to be selected by the structural and chemical restrictions on the amino acid at the P1 and P1' positions because these two amino acids have to bind perfectly at the active site for hydrolysis of the first peptide bond to occur. The hexameric assembly results in the merger of the open ends of the six substrate-binding channels from the LAP-A monomers to form a spacious central cavity allowing the hexameric LAP-A enzyme to simultaneously hydrolyze six peptides containing up to six amino acids each. The hexameric LAP-A enzyme may also hydrolyze long peptides or proteins if only one such substrate is bound to the hexamer because the substrate can extend through the central cavity and the two major solvent channels between the two LAP-A trimers.
- Department of Biochemistry, University of California-Riverside, Riverside, CA 92521, USA.
Organizational Affiliation: