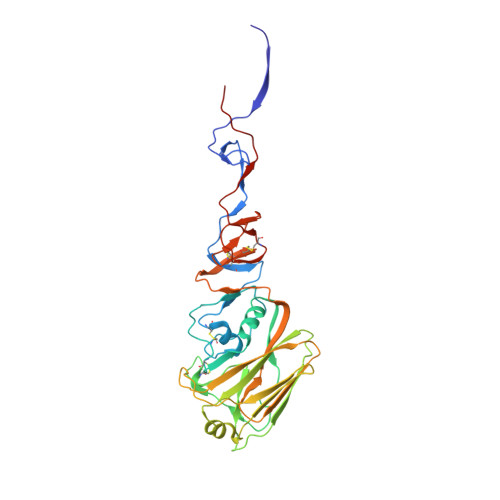
Structure and Receptor Binding Specificity of Hemagglutinin H13 from Avian Influenza A Virus H13N6
Lu, X., Qi, J., Shi, Y., Wang, M., Smith, D.F., Heimburg-Molinaro, J., Zhang, Y., Paulson, J.C., Xiao, H., Gao, G.F.(2013) J Virol 87: 9077-9085
- PubMed: 23760233 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.00235-13
- Primary Citation Related Structures:
4KPQ, 4KPS - PubMed Abstract:
Interspecies transmission (host switching/jumping) of influenza viruses is a key scientific question that must be addressed. In addition to the vigorous research on highly pathogenic avian influenza viruses (HPAIVs), studies of the mechanism of interspecies transmission of low-pathogenic avian influenza viruses (LPAIVs) could also provide insights into host tropism and virulence evolution. Influenza A viruses harboring hemagglutinin (HA) H13 (e.g., H13N6) are LPAIVs. In this study, soluble H13 HA glycoprotein was purified, and its receptor binding activity was characterized. The results revealed that H13 exclusively binds the avian α2-3-linked sialic acid receptor; no binding to the mammalian α2-6-linked sialic acid receptor was detected. Furthermore, the molecular basis of the H13 receptor binding specificity was revealed by comparative analysis of the crystal structures of both receptor-bound H13 and H5 HAs, which might be contributed by the hydrophobic residue V186. Work with an H13N186 mutant confirmed the importance of V186 in the receptor binding specificity of H13 HA, which shows that the mutant protein reduced the binding of an avian receptor analog but increased the binding of a human receptor analog. Detailed structural analysis also demonstrated that the conserved binding sites of the recently well-studied broadly neutralizing human monoclonal antibodies targeting the HA2 domain are found in H13. Our results expand our understanding of virulence evolution, receptor binding preference, and species tropism of the LPAIVs and HPAIVs.
- College of Veterinary Medicine, China Agricultural University, Beijing, China.
Organizational Affiliation: