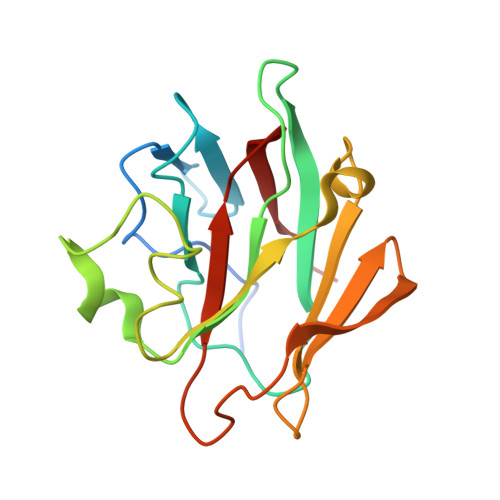
Crystal structure, biochemical and biophysical characterisation of NHR1 domain of E3 Ubiquitin ligase neutralized
Gupta, D., Beaufils, S., Vie, V., Paboeuf, G., Broadhurst, B., Schweisguth, F., Blundell, T.L., Bolanos-Garcia, V.M.(2013) Adv Enzyme Res 1: 61-75