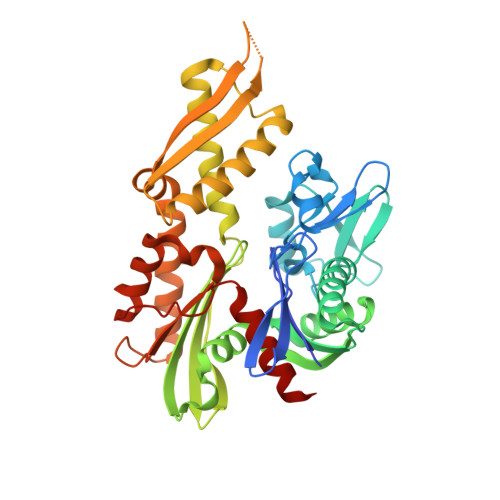
Crystal structure of the nucleotide-binding domain of mortalin, the mitochondrial Hsp70 chaperone.
Amick, J., Schlanger, S.E., Wachnowsky, C., Moseng, M.A., Emerson, C.C., Dare, M., Luo, W.I., Ithychanda, S.S., Nix, J.C., Cowan, J.A., Page, R.C., Misra, S.(2014) Protein Sci 23: 833-842
- PubMed: 24687350 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2466
- Primary Citation Related Structures:
4KBO - PubMed Abstract:
Mortalin, a member of the Hsp70-family of molecular chaperones, functions in a variety of processes including mitochondrial protein import and quality control, Fe-S cluster protein biogenesis, mitochondrial homeostasis, and regulation of p53. Mortalin is implicated in regulation of apoptosis, cell stress response, neurodegeneration, and cancer and is a target of the antitumor compound MKT-077. Like other Hsp70-family members, Mortalin consists of a nucleotide-binding domain (NBD) and a substrate-binding domain. We determined the crystal structure of the NBD of human Mortalin at 2.8 Å resolution. Although the Mortalin nucleotide-binding pocket is highly conserved relative to other Hsp70 family members, we find that its nucleotide affinity is weaker than that of Hsc70. A Parkinson's disease-associated mutation is located on the Mortalin-NBD surface and may contribute to Mortalin aggregation. We present structure-based models for how the Mortalin-NBD may interact with the nucleotide exchange factor GrpEL1, with p53, and with MKT-077. Our structure may contribute to the understanding of disease-associated Mortalin mutations and to improved Mortalin-targeting antitumor compounds.
- Department of Molecular Cardiology, The Cleveland Clinic, Cleveland, Ohio, 44195.
Organizational Affiliation: