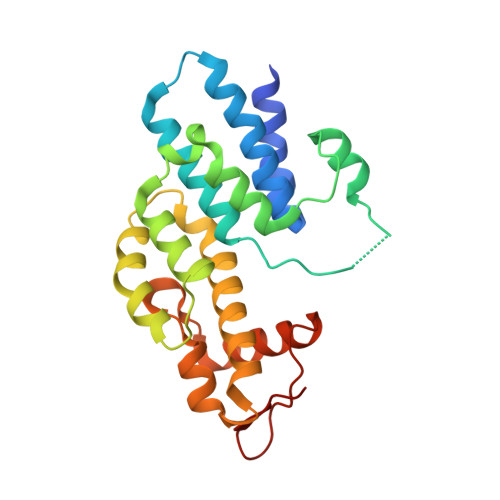
Structural integrity of the PCI domain of eIF3a/TIF32 is required for mRNA recruitment to the 43S pre-initiation complexes.
Khoshnevis, S., Gunisova, S., Vlckova, V., Kouba, T., Neumann, P., Beznoskova, P., Ficner, R., Valasek, L.S.(2014) Nucleic Acids Res 42: 4123-4139
- PubMed: 24423867 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkt1369
- Primary Citation Related Structures:
4K51 - PubMed Abstract:
Transfer of genetic information from genes into proteins is mediated by messenger RNA (mRNA) that must be first recruited to ribosomal pre-initiation complexes (PICs) by a mechanism that is still poorly understood. Recent studies showed that besides eIF4F and poly(A)-binding protein, eIF3 also plays a critical role in this process, yet the molecular mechanism of its action is unknown. We showed previously that the PCI domain of the eIF3c/NIP1 subunit of yeast eIF3 is involved in RNA binding. To assess the role of the second PCI domain of eIF3 present in eIF3a/TIF32, we performed its mutational analysis and identified a 10-Ala-substitution (Box37) that severely reduces amounts of model mRNA in the 43-48S PICs in vivo as the major, if not the only, detectable defect. Crystal structure analysis of the a/TIF32-PCI domain at 2.65-Å resolution showed that it is required for integrity of the eIF3 core and, similarly to the c/NIP1-PCI, is capable of RNA binding. The putative RNA-binding surface defined by positively charged areas contains two Box37 residues, R363 and K364. Their substitutions with alanines severely impair the mRNA recruitment step in vivo suggesting that a/TIF32-PCI represents one of the key domains ensuring stable and efficient mRNA delivery to the PICs.
- Department of Molecular Structural Biology, Institute for Microbiology and Genetics, George-August University, Goettingen, Germany, 37077 and Laboratory of Regulation of Gene Expression, Institute of Microbiology ASCR, Videnska 1083, 142 20 Prague, the Czech Republic.
Organizational Affiliation: