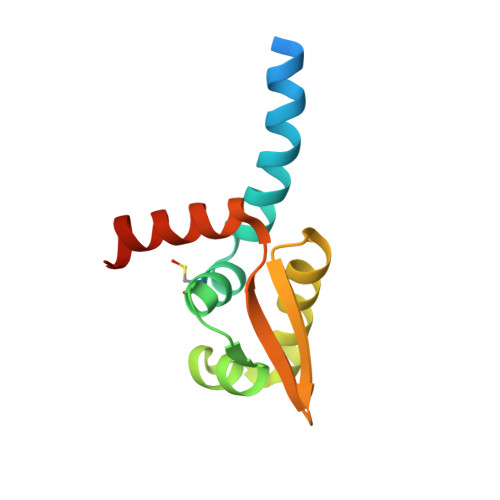
Crystal structure of HlyU, the hemolysin gene transcription activator, from Vibrio cholerae N16961 and functional implications.
Mukherjee, D., Datta, A.B., Chakrabarti, P.(2014) Biochim Biophys Acta 1844: 2346-2354
- PubMed: 25450504
- DOI: https://doi.org/10.1016/j.bbapap.2014.09.020
- Primary Citation of Related Structures:
4K2E, 4OOI - PubMed Abstract:
HlyU in Vibrio cholerae is known to be the transcriptional activator of the hemolysin gene, HlyA and possibly a regulator of other virulence factors influencing growth, colonization and pathogenicity of this infective agent. Here we report the crystal structure of HlyU from V. cholerae N16961 (HlyU_Vc) at 1.8Å. The protein, with five α-helices and three β-strands in the topology of α1-α2-β1-α3-α4-β2-β3-α5, forms a homodimer. Helices α3-α4 and a β sheet form the winged helix-turn-helix (wHTH) DNA-binding motif common to the transcription regulators of the SmtB/ArsR family. In spite of an overall fold similar to SmtB/ArsR family, it lacks any metal binding site seen in SmtB. A comparison of the dimeric interfaces showed that the one in SmtB is much larger and have salt bridges that can be disrupted to accommodate metal ions. A model of HlyU-DNA complex suggests bending of the DNA. Cys38 in the structure was found to be modified as sulfenic acid; the oxidized form was not seen in another structure solved under reducing condition. Although devoid of any metal binding site, the presence of a Cys residue exhibiting oxidation-reduction suggests the possibility of the existence of a redox switch in transcription regulation. A structure-based phylogenetic analysis of wHTH proteins revealed the segregation of metal and non-metal binding proteins as well as those in the latter group that are under redox control.
- Bioinformatics Centre, Bose Institute, P1/12 CIT Scheme VIIM, Kolkata 700054, India.
Organizational Affiliation: