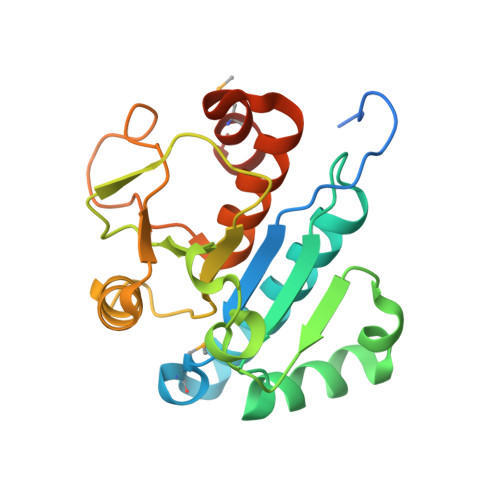
Crystal structure of tRNA m1G9 methyltransferase Trm10: insight into the catalytic mechanism and recognition of tRNA substrate.
Shao, Z., Yan, W., Peng, J., Zuo, X., Zou, Y., Li, F., Gong, D., Ma, R., Wu, J., Shi, Y., Zhang, Z., Teng, M., Li, X., Gong, Q.(2014) Nucleic Acids Res 42: 509-525
- PubMed: 24081582
- DOI: https://doi.org/10.1093/nar/gkt869
- Primary Citation of Related Structures:
4JWF, 4JWG, 4JWH, 4JWJ - PubMed Abstract:
Transfer RNA (tRNA) methylation is necessary for the proper biological function of tRNA. The N(1) methylation of guanine at Position 9 (m(1)G9) of tRNA, which is widely identified in eukaryotes and archaea, was found to be catalyzed by the Trm10 family of methyltransferases (MTases). Here, we report the first crystal structures of the tRNA MTase spTrm10 from Schizosaccharomyces pombe in the presence and absence of its methyl donor product S-adenosyl-homocysteine (SAH) and its ortholog scTrm10 from Saccharomyces cerevisiae in complex with SAH. Our crystal structures indicated that the MTase domain (the catalytic domain) of the Trm10 family displays a typical SpoU-TrmD (SPOUT) fold. Furthermore, small angle X-ray scattering analysis reveals that Trm10 behaves as a monomer in solution, whereas other members of the SPOUT superfamily all function as homodimers. We also performed tRNA MTase assays and isothermal titration calorimetry experiments to investigate the catalytic mechanism of Trm10 in vitro. In combination with mutational analysis and electrophoretic mobility shift assays, our results provide insights into the substrate tRNA recognition mechanism of Trm10 family MTases.
- Hefei National Laboratory for Physical Sciences at the Microscale and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui, 230026, People's Republic of China and X-ray Science Division, Advanced Photon Source, Argonne National Laboratory, Argonne, IL 60349, USA.
Organizational Affiliation: