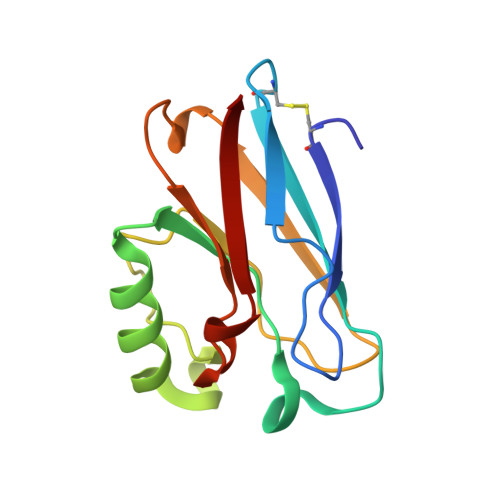
Mercury metallation of the copper protein azurin and structural insight into possible heavy metal reactivity.
Zampino, A.P., Masters, F.M., Bladholm, E.L., Panzner, M.J., Berry, S.M., Leeper, T.C., Ziegler, C.J.(2014) J Inorg Biochem 141: 152-160
- PubMed: 25265377 Search on PubMed
- DOI: https://doi.org/10.1016/j.jinorgbio.2014.09.003
- Primary Citation Related Structures:
4JKN - PubMed Abstract:
Mercury(II) metallation of Pseudomonas aeruginosa azurin has been characterized structurally and biochemically. The X-ray crystal structure at 1.5Å of mercury(II) metallated azurin confirms the coordination of mercury at the copper binding active site and a second surface site. These findings are further validated by NMR, Matrix-assisted laser desorption/ionization spectrometry (MALDI), and UV-visible spectroscopic methods indicating copper displacement from the wild-type protein. Bioinformatic analysis has identified homologous human protein domains computationally, and compared them to the structure of azurin, providing a model for human mercury interactions. Study of the mercury-azurin adduct, in combination with other known examples of protein-heavy metal interactions, could provide further insight into the chemical mechanisms of toxicological interactions, leading toward a global understanding of the biological speciation of toxic heavy metals.
- Department of Chemistry, The University of Akron, Akron, OH 44325, USA.
Organizational Affiliation: