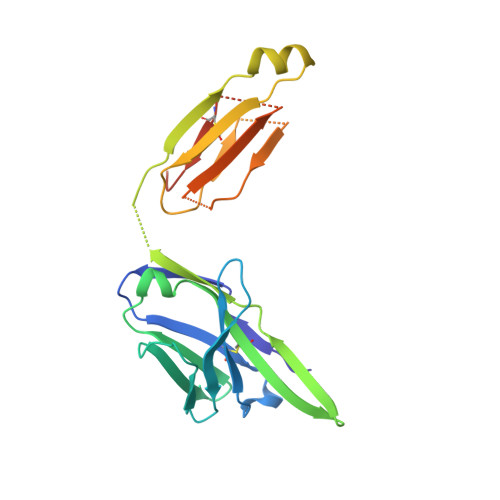
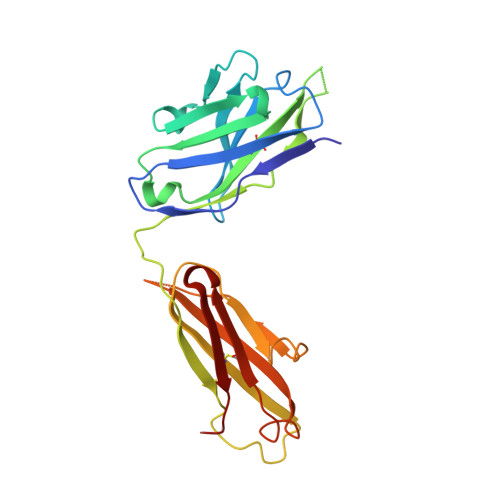
Linear and extended: a common polyglutamine conformation recognized by the three antibodies MW1, 1C2 and 3B5H10.
Klein, F.A., Zeder-Lutz, G., Cousido-Siah, A., Mitschler, A., Katz, A., Eberling, P., Mandel, J.L., Podjarny, A., Trottier, Y.(2013) Hum Mol Genet 22: 4215-4223
- PubMed: 23777629 Search on PubMed
- DOI: https://doi.org/10.1093/hmg/ddt273
- Primary Citation Related Structures:
4ISV, 4JJ5 - PubMed Abstract:
A long-standing pathomechanistic model proposes that the polyglutamine (polyQ)-length-dependent toxicity threshold observed in all polyQ diseases is triggered by a conformational change within the monomer that occurs only above a certain polyQ length. If true, this yet undefined and elusive mutant-specific toxic conformation would constitute a direct therapeutic target. Three anti-polyQ antibodies-MW1, 1C2 and 3B5H10-have been extensively used to probe the conformation of polyQ. The crystal structure of the MW1 epitope reveals a linear, non-pathogenic polyQ. In contrast, although the detailed structure of its epitope is unknown, the 3B5H10 antibody is widely advertised and used as a conformational antibody that recognizes the toxic conformation of expanded polyQ. We solved the crystal structure of the 1C2 antigen-binding domain (1C2-Fab) and performed a direct comparison between the 1C2, MW1 and 3B5H10 structures. The MW1 and 1C2 antibodies have similar sequences and structures, consistent with their binding to short polyQ and their polyQ length-discrimination properties. Unexpectedly, the 3B5H10 antibody also shares striking features with MW1 and 1C2, which prompted us to revisit its binding properties. We show that the 3B5H10 epitope is actually a short, non-pathogenic polyQ. All three antibodies MW1, 1C2 and 3B5H10 interact similarly with polyQ of various lengths, and bind small polyQ epitopes in similar linear and extended conformations. Together with studies published during the recent years, our work argues against the hypothesis that a mutant-specific conformation in monomeric polyQ molecules is the toxic entity responsible for polyQ diseases.
- Present address: Computational Chemistry and Biology Group-DETEMA, Facultad de Química, UdelaR, Isidoro de María 1620 piso 3, CC1157, Montevideo, Uruguay.
Organizational Affiliation: