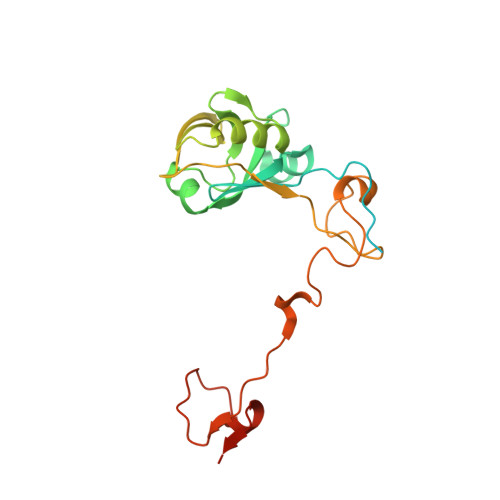
Structural insight into magnetochrome-mediated magnetite biomineralization.
Siponen, M.I., Legrand, P., Widdrat, M., Jones, S.R., Zhang, W.J., Chang, M.C., Faivre, D., Arnoux, P., Pignol, D.(2013) Nature 502: 681-684
- PubMed: 24097349 Search on PubMed
- DOI: https://doi.org/10.1038/nature12573
- Primary Citation Related Structures:
4JJ0, 4JJ3 - PubMed Abstract:
Magnetotactic bacteria align along the Earth's magnetic field using an organelle called the magnetosome, a biomineralized magnetite (Fe(II)Fe(III)2O4) or greigite (Fe(II)Fe(III)2S4) crystal embedded in a lipid vesicle. Although the need for both iron(II) and iron(III) is clear, little is known about the biological mechanisms controlling their ratio. Here we present the structure of the magnetosome-associated protein MamP and find that it is built on a unique arrangement of a self-plugged PDZ domain fused to two magnetochrome domains, defining a new class of c-type cytochrome exclusively found in magnetotactic bacteria. Mutational analysis, enzyme kinetics, co-crystallization with iron(II) and an in vitro MamP-assisted magnetite production assay establish MamP as an iron oxidase that contributes to the formation of iron(III) ferrihydrite eventually required for magnetite crystal growth in vivo. These results demonstrate the molecular mechanisms of iron management taking place inside the magnetosome and highlight the role of magnetochrome in iron biomineralization.
- 1] Commissariat à l'Energie Atomique et aux Energies Alternatives, Direction des Sciences du Vivant, Institut de Biologie Environnementale et de Biotechnologies,, F-13108, France [2] Centre National de la Recherche Scientifique, Unité Mixte de Recherche Biologie Végétale et Microbiologie Environnementales, Saint-Paul-lez-Durance, F-13108, France [3] Aix-Marseille Université, Saint-Paul-lez-Durance, F-13108, France.
Organizational Affiliation: