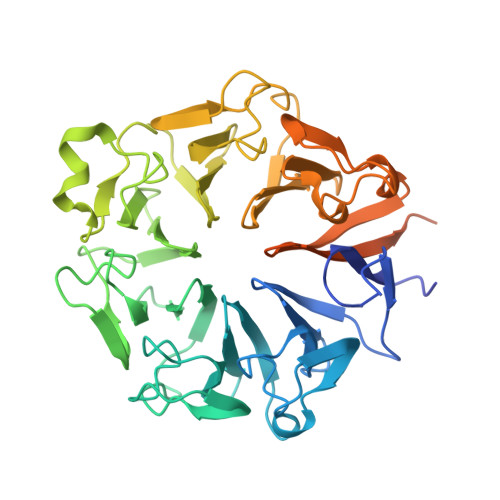
The interplay between RPGR, PDE-delta and Arl2/3 regulate the ciliary targeting of farnesylated cargo.
Watzlich, D., Vetter, I., Gotthardt, K., Miertzschke, M., Chen, Y.X., Wittinghofer, A., Ismail, S.(2013) EMBO Rep 14: 465-472
- PubMed: 23559067
- DOI: https://doi.org/10.1038/embor.2013.37
- Primary Citation Related Structures:
4JHN, 4JHP - PubMed Abstract:
Defects in primary cilia result in human diseases known as ciliopathies. The retinitis pigmentosa GTPase regulator (RPGR), mutated in the most severe form of the eye disease, is located at the transition zone of the ciliary organelle. The RPGR-interacting partner PDEδ is involved in trafficking of farnesylated ciliary cargo, but the significance of this interaction is unknown. The crystal structure of the propeller domain of RPGR shows the location of patient mutations and how they perturb the structure. The RPGR·PDEδ complex structure shows PDEδ on a highly conserved surface patch of RPGR. Biochemical experiments and structural considerations show that RPGR can bind with high affinity to cargo-loaded PDEδ and exposes the Arl2/Arl3-binding site on PDEδ. On the basis of these results, we propose a model where RPGR is acting as a scaffold protein recruiting cargo-loaded PDEδ and Arl3 to release lipidated cargo into cilia.
- Structural Biology Group, Max Planck Institute for Molecular Physiology, Otto-Hahn-Strasse 11, Dortmund 44227, Germany.
Organizational Affiliation: