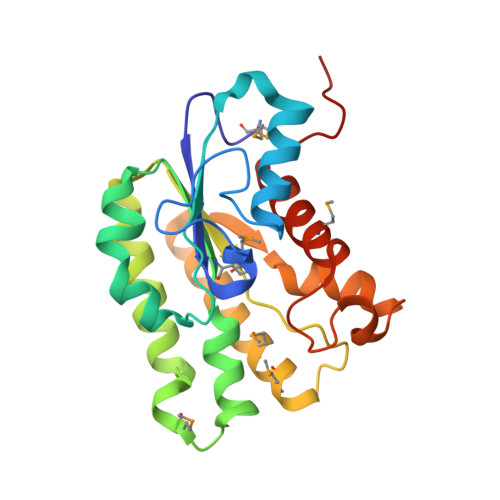
A unique octameric structure of Axe2, an intracellular acetyl-xylooligosaccharide esterase from Geobacillus stearothermophilus.
Lansky, S., Alalouf, O., Solomon, H.V., Alhassid, A., Govada, L., Chayen, N.E., Belrhali, H., Shoham, Y., Shoham, G.(2014) Acta Crystallogr D Biol Crystallogr 70: 261-278
- PubMed: 24531461 Search on PubMed
- DOI: https://doi.org/10.1107/S139900471302840X
- Primary Citation Related Structures:
3W7V, 4JHL, 4JKO - PubMed Abstract:
Geobacillus stearothermophilus T6 is a thermophilic, Gram-positive soil bacterium that possesses an extensive and highly regulated hemicellulolytic system, allowing the bacterium to efficiently degrade high-molecular-weight polysaccharides such as xylan, arabinan and galactan. As part of the xylan-degradation system, the bacterium uses a number of side-chain-cleaving enzymes, one of which is Axe2, a 219-amino-acid intracellular serine acetylxylan esterase that removes acetyl side groups from xylooligosaccharides. Bioinformatic analyses suggest that Axe2 belongs to the lipase GDSL family and represents a new family of carbohydrate esterases. In the current study, the detailed three-dimensional structure of Axe2 is reported, as determined by X-ray crystallography. The structure of the selenomethionine derivative Axe2-Se was initially determined by single-wavelength anomalous diffraction techniques at 1.70 Å resolution and was used for the structure determination of wild-type Axe2 (Axe2-WT) and the catalytic mutant Axe2-S15A at 1.85 and 1.90 Å resolution, respectively. These structures demonstrate that the three-dimensional structure of the Axe2 monomer generally corresponds to the SGNH hydrolase fold, consisting of five central parallel β-sheets flanked by two layers of helices (eight α-helices and five 310-helices). The catalytic triad residues, Ser15, His194 and Asp191, are lined up along a substrate channel situated on the concave surface of the monomer. Interestingly, the Axe2 monomers are assembled as a `doughnut-shaped' homo-octamer, presenting a unique quaternary structure built of two staggered tetrameric rings. The eight active sites are organized in four closely situated pairs, which face the relatively wide internal cavity. The biological relevance of this octameric structure is supported by independent results obtained from gel-filtration, TEM and SAXS experiments. These data and their comparison to the structural data of related hydrolases are used for a more general discussion focusing on the structure-function relationships of enzymes of this category.
- Institute of Chemistry and the Laboratory for Structural Chemistry and Biology, The Hebrew University of Jerusalem, 91904 Jerusalem, Israel.
Organizational Affiliation: