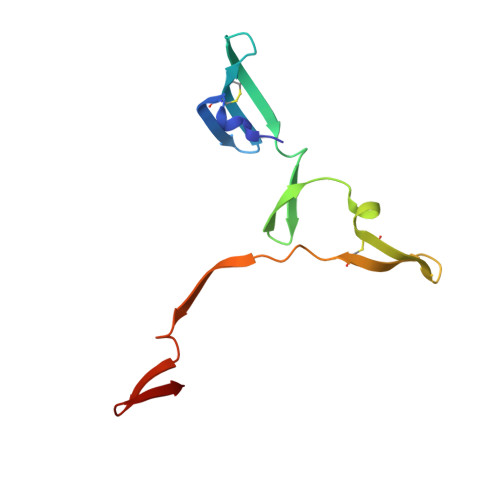
Different 3D domain-swapped oligomeric cyanovirin-N structures suggest trapped folding intermediates.
Koharudin, L.M., Liu, L., Gronenborn, A.M.(2013) Proc Natl Acad Sci U S A 110: 7702-7707
- PubMed: 23610431
- DOI: https://doi.org/10.1073/pnas.1300327110
- Primary Citation of Related Structures:
4J4C, 4J4D, 4J4E, 4J4F, 4J4G - PubMed Abstract:
Although it has long been established that the amino acid sequence encodes the fold of a protein, how individual proteins arrive at their final conformation is still difficult to predict, especially for oligomeric structures. Here, we present a comprehensive characterization of oligomeric species of cyanovirin-N that all are formed by a polypeptide chain with the identical amino acid sequence. Structures of the oligomers were determined by X-ray crystallography, and each one exhibits 3D domain swapping. One unique 3D domain-swapped structure is observed for the trimer, while for both dimer and tetramer, two different 3D domain-swapped structures were obtained. In addition to the previously identified hinge-loop region of the 3D domain-swapped dimer, which resides between strands β5 and β6 in the middle of the polypeptide sequence, another hinge-loop region is observed between strands β7 and β8 in the structures. Plasticity in these two regions allows for variability in dihedral angles and concomitant differences in chain conformation that results in the differently 3D domain-swapped multimers. Based on all of the different structures, we propose possible folding pathways for this protein. Altogether, our results illuminate the amazing ability of cyanovirin-N to proceed down different folding paths and provide general insights into oligomer formation via 3D domain swapping.
- Department of Structural Biology, University of Pittsburgh School of Medicine, Pittsburgh, PA 15260, USA.
Organizational Affiliation: