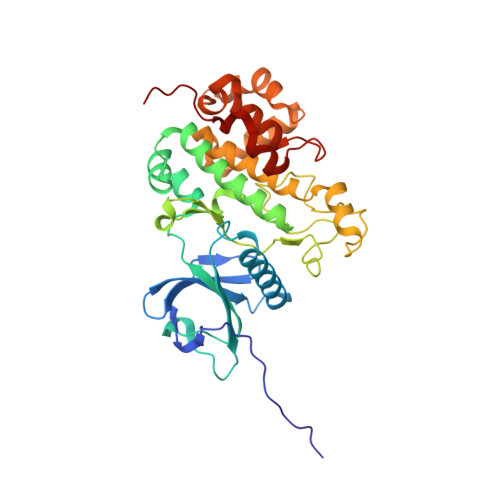
Structure of the catalytic domain of a state transition kinase homolog from Micromonas algae
Guo, J., Wei, X., Li, M., Pan, X., Chang, W., Liu, Z.(2013) Protein Cell 4: 607-619
- PubMed: 23794031 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s13238-013-3034-9
- Primary Citation Related Structures:
4IX3, 4IX4, 4IX5, 4IX6 - PubMed Abstract:
Under natural environments, plants and algae have evolved various photosynthetic acclimation mechanisms in response to the constantly changing light conditions. The state transition and long-term response processes in photosynthetic acclimation involve remodeling and composition alteration of thylakoid membrane. A chloroplast protein kinase named Stt7/STN7 has been found to have pivotal roles in both state transition and long-term response. Here we report the crystal structures of the kinase domain of a putative Stt7/STN7 homolog from Micromonas sp. RCC299 (MsStt7d) in the apo form and in complex with various nucleotide substrates. MsStt7d adopts a canonical protein kinase fold and contains all the essential residues at the active site. A novel hairpin motif, found to be a conserved feature of the Stt7/STN7 family and indispensable for the kinase stability, interacts with the activation loop and fixes it in an active conformation. We have also demonstrated that MsStt7d is a dualspecifi city kinase that phosphorylates both Thr and Tyr residues. Moreover, preliminary in vitro data suggest that it might be capable of phosphorylating a consensus N-terminal pentapeptide of light-harvesting proteins Micromonas Lhcp4 and Arabidopsis Lhcb1 directly. The potential peptide/protein substrate binding site is predicted based on the location of a pseudo-substrate contributed by the adjacent molecule within the crystallographic dimer. The structural and biochemical data presented here provide a framework for an improved understanding on the role of Stt7/STN7 in photosynthetic acclimation.
- National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, China.
Organizational Affiliation: