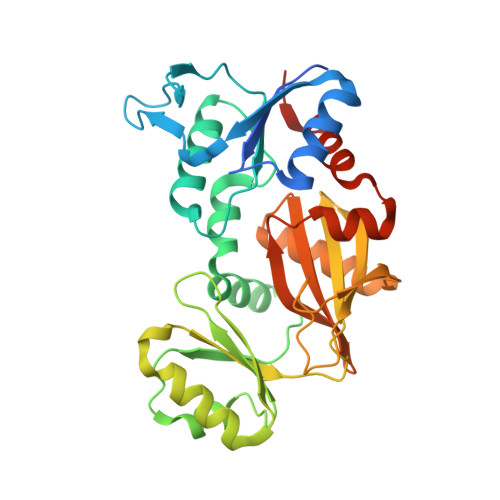
Structure and function of Escherichia coli RimK, an ATP-grasp fold, l-glutamyl ligase enzyme.
Zhao, G., Jin, Z., Wang, Y., Allewell, N.M., Tuchman, M., Shi, D.(2013) Proteins 81: 1847-1854
- PubMed: 23609986 Search on PubMed
- DOI: https://doi.org/10.1002/prot.24311
- Primary Citation Related Structures:
4IWX, 4IWY - PubMed Abstract:
We report herein the crystal structure of Escherichia coli RimK at a resolution of 2.85 Å, an enzyme that catalyzes the post-translational addition of up to 15 C-terminal glutamate residues to ribosomal protein S6. The structure belongs to the ATP-grasp superfamily and is organized as a tetramer, consistent with gel filtration analysis. Each subunit consists of three distinct structural domains and the active site is located in the cleft between these domains. The catalytic reaction appears to occur at the junction between the three domains as ATP binds between the B and C domains, and other substrates bind nearby.
- Department of Integrative Systems Biology, Center for Genetic Medicine Research, Children's National Medical Center, The George Washington University, Washington, DC, 20010.
Organizational Affiliation: