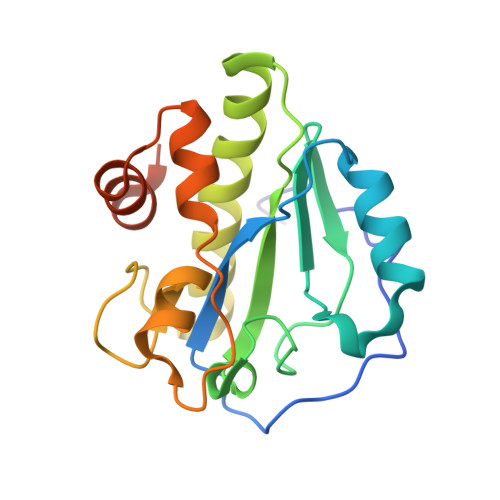
Structural and functional insights into peptidoglycan access for the lytic amidase LytA of Streptococcus pneumoniae.
Mellroth, P., Sandalova, T., Kikhney, A., Vilaplana, F., Hesek, D., Lee, M., Mobashery, S., Normark, S., Svergun, D., Henriques-Normark, B., Achour, A.(2014) mBio 5: e01120-e01113
- PubMed: 24520066 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mBio.01120-13
- Primary Citation Related Structures:
4IVV, 4IWT - PubMed Abstract:
The cytosolic N-acetylmuramoyl-l-alanine amidase LytA protein of Streptococcus pneumoniae, which is released by bacterial lysis, associates with the cell wall via its choline-binding motif. During exponential growth, LytA accesses its peptidoglycan substrate to cause lysis only when nascent peptidoglycan synthesis is stalled by nutrient starvation or β-lactam antibiotics. Here we present three-dimensional structures of LytA and establish the requirements for substrate binding and catalytic activity. The solution structure of the full-length LytA dimer reveals a peculiar fold, with the choline-binding domains forming a rigid V-shaped scaffold and the relatively more flexible amidase domains attached in a trans position. The 1.05-Å crystal structure of the amidase domain reveals a prominent Y-shaped binding crevice composed of three contiguous subregions, with a zinc-containing active site localized at the bottom of the branch point. Site-directed mutagenesis was employed to identify catalytic residues and to investigate the relative impact of potential substrate-interacting residues lining the binding crevice for the lytic activity of LytA. In vitro activity assays using defined muropeptide substrates reveal that LytA utilizes a large substrate recognition interface and requires large muropeptide substrates with several connected saccharides that interact with all subregions of the binding crevice for catalysis. We hypothesize that the substrate requirements restrict LytA to the sites on the cell wall where nascent peptidoglycan synthesis occurs. Streptococcus pneumoniae is a human respiratory tract pathogen responsible for millions of deaths annually. Its major pneumococcal autolysin, LytA, is required for autolysis and fratricidal lysis and functions as a virulence factor that facilitates the spread of toxins and factors involved in immune evasion. LytA is also activated by penicillin and vancomycin and is responsible for the lysis induced by these antibiotics. The factors that regulate the lytic activity of LytA are unclear, but it was recently demonstrated that control is at the level of substrate recognition and that LytA required access to the nascent peptidoglycan. The present study was undertaken to structurally and functionally investigate LytA and its substrate-interacting interface and to determine the requirements for substrate recognition and catalysis. Our results reveal that the amidase domain comprises a complex substrate-binding crevice and needs to interact with a large-motif epitope of peptidoglycan for catalysis.