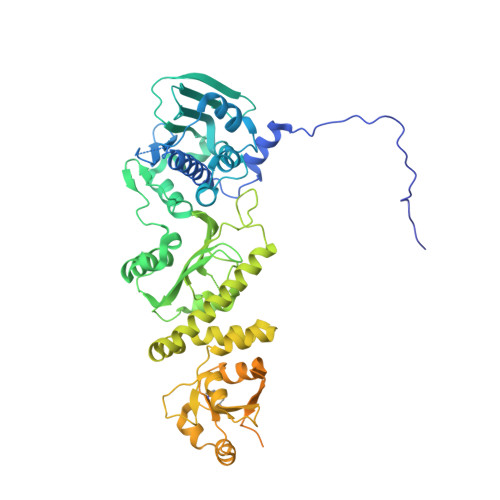
Structural asymmetry in the closed state of mitochondrial Hsp90 (TRAP1) supports a two-step ATP hydrolysis mechanism.
Lavery, L.A., Partridge, J.R., Ramelot, T.A., Elnatan, D., Kennedy, M.A., Agard, D.A.(2014) Mol Cell 53: 330-343
- PubMed: 24462206 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2013.12.023
- Primary Citation Related Structures:
4IPE, 4IVG, 4IYN, 4J0B - PubMed Abstract:
While structural symmetry is a prevailing feature of homo-oligomeric proteins, asymmetry provides unique mechanistic opportunities. We present the crystal structure of full-length TRAP1, the mitochondrial Hsp90 molecular chaperone, in a catalytically active closed state. The TRAP1 homodimer adopts a distinct, asymmetric conformation, where one protomer is reconfigured via a helix swap at the middle:C-terminal domain (MD:CTD) interface. This interface plays a critical role in client binding. Solution methods validate the asymmetry and show extension to Hsp90 homologs. Point mutations that disrupt unique contacts at each MD:CTD interface reduce catalytic activity and substrate binding and demonstrate that each protomer needs access to both conformations. Crystallographic data on a dimeric NTD:MD fragment suggests that asymmetry arises from strain induced by simultaneous NTD and CTD dimerization. The observed asymmetry provides the potential for an additional step in the ATPase cycle, allowing sequential ATP hydrolysis steps to drive both client remodeling and client release.
- Howard Hughes Medical Institute and the Department of Biochemistry and Biophysics, University of California, San Francisco, San Francisco, CA 94158, USA.
Organizational Affiliation: