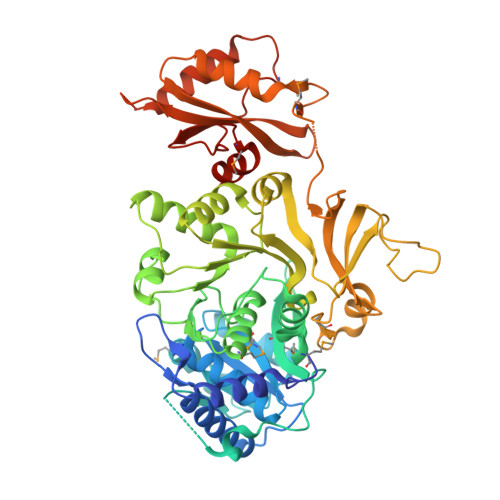
Structures of Mycobacterium tuberculosis FadD10 protein reveal a new type of adenylate-forming enzyme.
Liu, Z., Ioerger, T.R., Wang, F., Sacchettini, J.C.(2013) J Biological Chem 288: 18473-18483
- PubMed: 23625916 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.466912
- Primary Citation Related Structures:
4IR7, 4ISB - PubMed Abstract:
Mycobacterium tuberculosis has a group of 34 FadD proteins that belong to the adenylate-forming superfamily. They are classified as either fatty acyl-AMP ligases (FAALs) or fatty acyl-CoA ligases based on sequence analysis. FadD10, involved in the synthesis of a virulence-related lipopeptide, was mis-annotated as a fatty acyl-CoA ligase; however, it is in fact a FAAL that transfers fatty acids to an acyl carrier protein (Rv0100). In this study, we have determined the structures of FadD10 in both the apo-form and the complexed form with dodecanoyl-AMP, where we see for the first time an adenylate-forming enzyme that does not adopt a closed conformation for catalysis. Indeed, this novel conformation of FadD10, facilitated by its unique inter-domain and intermolecular interactions, is critical for the enzyme to carry out the acyl transfer onto Rv0100 rather than coenzyme A. This contradicts the existing model of FAALs that rely on an insertion motif for the acyltransferase specificity and thus makes FadD10 a new type of FAAL. We have also characterized the fatty acid preference of FadD10 through biological and structural analyses, and the data indicate long chain saturated fatty acids as the biological substrates of the enzyme.
- Department of Chemistry, Texas A&M University, College Station, Texas 77842-3012, USA.
Organizational Affiliation: