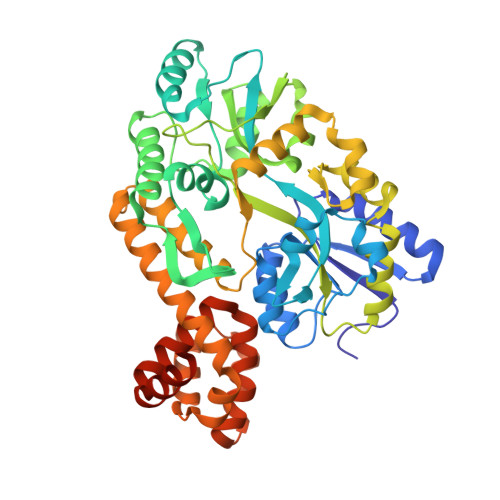
Structure of the caspase-recruitment domain from a zebrafish guanylate-binding protein.
Jin, T., Huang, M., Smith, P., Jiang, J., Xiao, T.S.(2013) Acta Crystallogr Sect F Struct Biol Cryst Commun 69: 855-860
- PubMed: 23908027 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309113015558
- Primary Citation Related Structures:
4IRL - PubMed Abstract:
The caspase-recruitment domain (CARD) mediates homotypic protein-protein interactions that assemble large oligomeric signaling complexes such as the inflammasomes during innate immune responses. Structural studies of the mammalian CARDs demonstrate that their six-helix bundle folds belong to the death-domain superfamily, whereas such studies have not been reported for other organisms. Here, the zebrafish interferon-induced guanylate-binding protein 1 (zIGBP1) was identified that contains an N-terminal GTPase domain and a helical domain typical of the mammalian guanylate-binding proteins, followed by a FIIND domain and a C-terminal CARD similar to the mammalian inflammasome proteins NLRP1 and CARD8. The structure of the zIGBP1 CARD as a fusion with maltose-binding protein was determined at 1.47 Å resolution. This revealed a six-helix bundle fold similar to the NLRP1 CARD structure with the bent α1 helix typical of all known CARD structures. The zIGBP1 CARD surface contains a positively charged patch near its α1 and α4 helices and a negatively charged patch near its α2, α3 and α5 helices, which may mediate its interaction with partner domains. Further studies using binding assays and other analyses will be required in order to address the physiological function(s) of this zebrafish protein.
- Structural Immunobiology Unit, Laboratory of Immunology, National Institute of Allergy and Infectious Diseases, National Institutes of Health, 4 Memorial Drive, Building 4, Room 228, Bethesda, MD 20892-0430, USA.
Organizational Affiliation: