Structural insights into the substrate specificity of a 6-phospho-&[beta]-glucosidase BglA-2 from Streptococcus pneumoniae TIGR4
Yu, W.L., Jiang, Y.L., Pikis, A., Cheng, W., Bai, X.H., Ren, Y.M., Thompson, J., Zhou, C.Z., Chen, Y.(2013) J Biological Chem 288: 14949-14958
- PubMed: 23580646 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.454751
- Primary Citation Related Structures:
4IPL, 4IPN - PubMed Abstract:
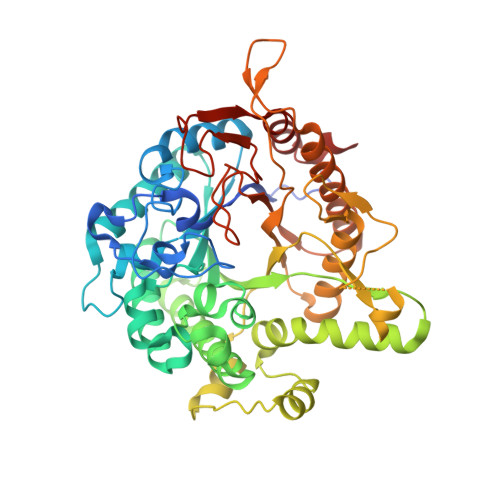
The 6-phospho-β-glucosidase BglA-2 (EC 3.2.1.86) from glycoside hydrolase family 1 (GH-1) catalyzes the hydrolysis of β-1,4-linked cellobiose 6-phosphate (cellobiose-6'P) to yield glucose and glucose 6-phosphate. Both reaction products are further metabolized by the energy-generating glycolytic pathway. Here, we present the first crystal structures of the apo and complex forms of BglA-2 with thiocellobiose-6'P (a non-metabolizable analog of cellobiose-6'P) at 2.0 and 2.4 Å resolution, respectively. Similar to other GH-1 enzymes, the overall structure of BglA-2 from Streptococcus pneumoniae adopts a typical (β/α)8 TIM-barrel, with the active site located at the center of the convex surface of the β-barrel. Structural analyses, in combination with enzymatic data obtained from site-directed mutant proteins, suggest that three aromatic residues, Tyr(126), Tyr(303), and Trp(338), at subsite +1 of BglA-2 determine substrate specificity with respect to 1,4-linked 6-phospho-β-glucosides. Moreover, three additional residues, Ser(424), Lys(430), and Tyr(432) of BglA-2, were found to play important roles in the hydrolytic selectivity toward phosphorylated rather than non-phosphorylated compounds. Comparative structural analysis suggests that a tryptophan versus a methionine/alanine residue at subsite -1 may contribute to the catalytic and substrate selectivity with respect to structurally similar 6-phospho-β-galactosidases and 6-phospho-β-glucosidases assigned to the GH-1 family.
- Hefei National Laboratory for Physical Sciences at the Microscale and School of Life Sciences, University of Science and Technology of China, Hefei Anhui 230027, China.
Organizational Affiliation: