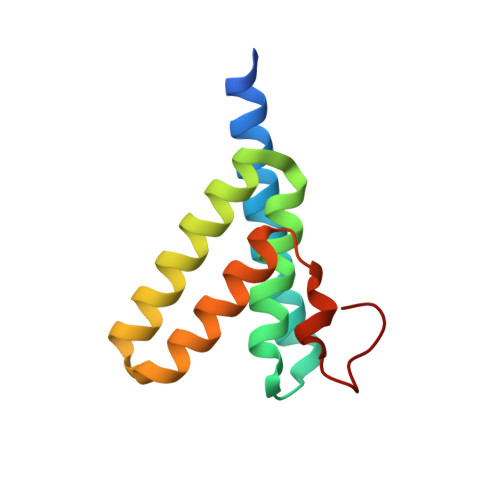
Structural mechanism of serum amyloid A-mediated inflammatory amyloidosis.
Lu, J., Yu, Y., Zhu, I., Cheng, Y., Sun, P.D.(2014) Proc Natl Acad Sci U S A 111: 5189-5194
- PubMed: 24706838 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1322357111
- Primary Citation Related Structures:
4IP8, 4IP9 - PubMed Abstract:
Serum amyloid A (SAA) represents an evolutionarily conserved family of inflammatory acute-phase proteins. It is also a major constituent of secondary amyloidosis. To understand its function and structural transition to amyloid, we determined a structure of human SAA1.1 in two crystal forms, representing a prototypic member of the family. Native SAA1.1 exists as a hexamer, with subunits displaying a unique four-helix bundle fold stabilized by its long C-terminal tail. Structure-based mutational studies revealed two positive-charge clusters, near the center and apex of the hexamer, that are involved in SAA association with heparin. The binding of high-density lipoprotein involves only the apex region of SAA and can be inhibited by heparin. Peptide amyloid formation assays identified the N-terminal helices 1 and 3 as amyloidogenic peptides of SAA1.1. Both peptides are secluded in the hexameric structure of SAA1.1, suggesting that the native SAA is nonpathogenic. Furthermore, dissociation of the SAA hexamer appears insufficient to initiate amyloidogenic transition, and proteolytic cleavage or removal of the C-terminal tail of SAA resulted in formation of various-sized structural aggregates containing ∼5-nm regular repeating protofibril-like units. The combined structural and functional studies provide mechanistic insights into the pathogenic contribution of glycosaminoglycan in SAA1.1-mediated AA amyloid formation.
- Structural Immunology Section, Laboratory of Immunogenetics, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Rockville, MD 20852.
Organizational Affiliation: