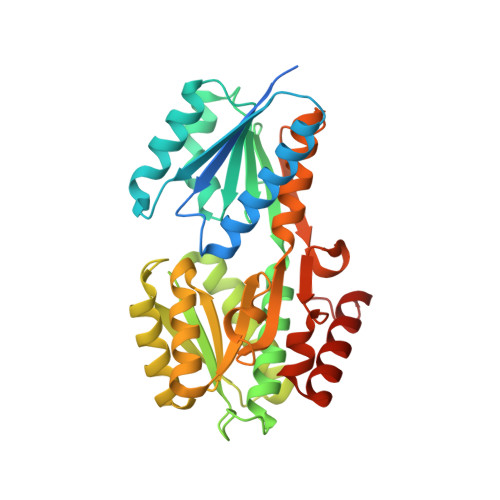
Evidence for an ABC-type riboflavin transporter system in pathogenic spirochetes.
Deka, R.K., Brautigam, C.A., Biddy, B.A., Liu, W.Z., Norgard, M.V.(2013) mBio 4: e00615-e00612
- PubMed: 23404400 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mBio.00615-12
- Primary Citation Related Structures:
4IIL - PubMed Abstract:
Bacterial transporter proteins are involved in the translocation of many essential nutrients and metabolites. However, many of these key bacterial transport systems remain to be identified, including those involved in the transport of riboflavin (vitamin B(2)). Pathogenic spirochetes lack riboflavin biosynthetic pathways, implying reliance on obtaining riboflavin from their hosts. Using structural and functional characterizations of possible ligand-binding components, we have identified an ABC-type riboflavin transport system within pathogenic spirochetes. The putative lipoprotein ligand-binding components of these systems from three different spirochetes were cloned, hyperexpressed in Escherichia coli, and purified to homogeneity. Solutions of all three of the purified recombinant proteins were bright yellow. UV-visible spectra demonstrated that these proteins were likely flavoproteins; electrospray ionization mass spectrometry and thin-layer chromatography confirmed that they contained riboflavin. A 1.3-Å crystal structure of the protein (TP0298) encoded by Treponema pallidum, the syphilis spirochete, demonstrated that the protein's fold is similar to the ligand-binding components of ABC-type transporters. The structure also revealed other salient details of the riboflavin binding site. Comparative bioinformatics analyses of spirochetal genomes, coupled with experimental validation, facilitated the discovery of this new ABC-type riboflavin transport system(s). We denote the ligand-binding component as riboflavin uptake transporter A (RfuA). Taken together, it appears that pathogenic spirochetes have evolved an ABC-type transport system (RfuABCD) for survival in their host environments, particularly that of the human host. Syphilis remains a public health problem, but very little is known about the causative bacterium. This is because Treponema pallidum still cannot be cultured in the laboratory. Rather, T. pallidum must be cultivated in laboratory rabbits, a restriction that poses many insurmountable experimental obstacles. Approaches to learn more about the structure and function of T. pallidum's cell envelope, which is both the physical and functional interface between T. pallidum and its human host, are severely limited. One approach for elucidating T. pallidum's cell envelope has been to determine the three-dimensional structures of its membrane lipoproteins, molecules that serve many critical survival functions. Herein, we describe a previously unknown transport system that T. pallidum uses to import riboflavin, an essential nutrient for the organism's survival. Moreover, we found that this transport system is present in other pathogenic spirochetes. This is the first description of this new type of bacterial riboflavin transport system.
- Department of Microbiology, University of Texas, Southwestern Medical Center, Dallas, Texas, USA.
Organizational Affiliation: