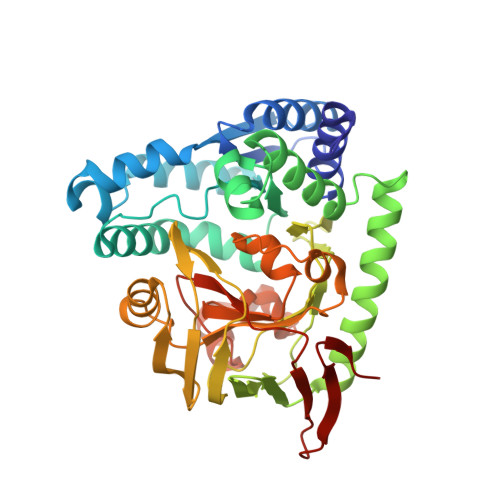
Structure-based elucidation of the regulatory mechanism for aminopeptidase activity.
Ta, H.M., Bae, S., Han, S., Song, J., Ahn, T.K., Hohng, S., Lee, S., Kim, K.K.(2013) Acta Crystallogr D Biol Crystallogr 69: 1738-1747
- PubMed: 23999297 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444913012651
- Primary Citation Related Structures:
4ICQ, 4ICR, 4ICS - PubMed Abstract:
The specificity of proteases for the residues in and length of substrates is key to understanding their regulatory mechanism, but little is known about length selectivity. Crystal structure analyses of the bacterial aminopeptidase PepS, combined with functional and single-molecule FRET assays, have elucidated a molecular basis for length selectivity. PepS exists in open and closed conformations. Substrates can access the binding hole in the open conformation, but catalytic competency is only achieved in the closed conformation by formation of the S1 binding pocket and proximal movement of Glu343, a general base, to the cleavage site. Hence, peptides longer than the depth of the binding hole block the transition from the open to the closed conformation, and thus length selection is a prerequisite for catalytic activation. A triple-sieve interlock mechanism is proposed featuring the coupling of length selectivity with residue specificity and active-site positioning.
- Department of Molecular Cell Biology, School of Medicine, Samsung Biomedical Research Institute, Sungkyunkwan University, Suwon 440-746, Republic of Korea.
Organizational Affiliation: