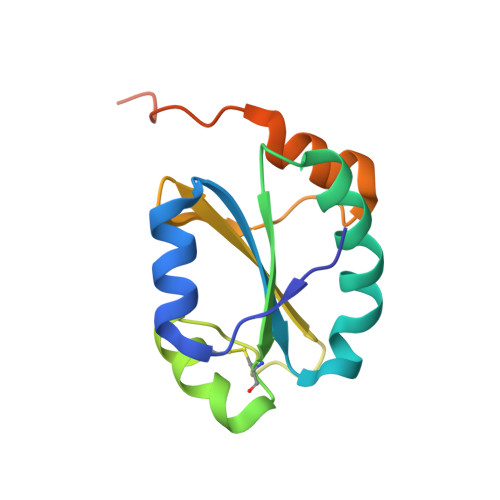
Structure of the Non-Catalytic Domain of the Protein Disulfide Isomerase-Related Protein (PDIR) Reveals Function in Protein Binding.
Vinaik, R., Kozlov, G., Gehring, K.(2013) PLoS One 8: e62021-e62021
- PubMed: 23614004 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0062021
- Primary Citation Related Structures:
4I6X - PubMed Abstract:
Protein disulfide isomerases comprise a large family of enzymes responsible for catalyzing the proper oxidation and folding of newly synthesized proteins in the endoplasmic reticulum (ER). Protein disulfide isomerase-related (PDIR) protein (also known as PDIA5) is a specialized member that participates in the folding of α1-antitrypsin and N-linked glycoproteins. Here, the crystal structure of the non-catalytic domain of PDIR was determined to 1.5 Å resolution. The structure adopts a thioredoxin-like fold stabilized by a structural disulfide bridge with a positively charged binding surface for interactions with the ER chaperones, calreticulin and ERp72. Crystal contacts between molecules potentially mimic the interactions of PDIR with misfolded substrate proteins. The results suggest that the non-catalytic domain of PDIR plays a key role in the recognition of protein partners and substrates.
- Department of Biochemistry, Groupe de Recherche Axé sur la Structure des Protéines, McGill University, Montréal, Québec, Canada.
Organizational Affiliation: