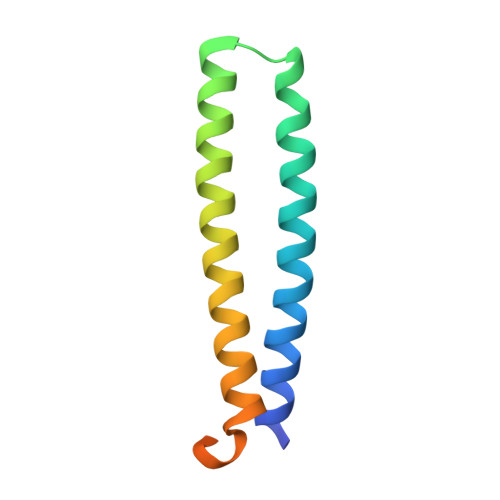
Heterologous expression of mycobacterial Esx complexes in Escherichia coli for structural studies is facilitated by the use of maltose binding protein fusions.
Arbing, M.A., Chan, S., Harris, L., Kuo, E., Zhou, T.T., Ahn, C.J., Nguyen, L., He, Q., Lu, J., Menchavez, P.T., Shin, A., Holton, T., Sawaya, M.R., Cascio, D., Eisenberg, D.(2013) PLoS One 8: e81753-e81753
- PubMed: 24312350
- DOI: https://doi.org/10.1371/journal.pone.0081753
- Primary Citation Related Structures:
3OGI, 3Q4H, 4GZR, 4I0X - PubMed Abstract:
The expression of heteroligomeric protein complexes for structural studies often requires a special coexpression strategy. The reason is that the solubility and proper folding of each subunit of the complex requires physical association with other subunits of the complex. The genomes of pathogenic mycobacteria encode many small protein complexes, implicated in bacterial fitness and pathogenicity, whose characterization may be further complicated by insolubility upon expression in Escherichia coli, the most common heterologous protein expression host. As protein fusions have been shown to dramatically affect the solubility of the proteins to which they are fused, we evaluated the ability of maltose binding protein fusions to produce mycobacterial Esx protein complexes. A single plasmid expression strategy using an N-terminal maltose binding protein fusion to the CFP-10 homolog proved effective in producing soluble Esx protein complexes, as determined by a small-scale expression and affinity purification screen, and coupled with intracellular proteolytic cleavage of the maltose binding protein moiety produced protein complexes of sufficient purity for structural studies. In comparison, the expression of complexes with hexahistidine affinity tags alone on the CFP-10 subunits failed to express in amounts sufficient for biochemical characterization. Using this strategy, six mycobacterial Esx complexes were expressed, purified to homogeneity, and subjected to crystallization screening and the crystal structures of the Mycobacterium abscessus EsxEF, M. smegmatis EsxGH, and M. tuberculosis EsxOP complexes were determined. Maltose binding protein fusions are thus an effective method for production of Esx complexes and this strategy may be applicable for production of other protein complexes.
- UCLA-DOE Institute for Genomics and Proteomics, University of California Los Angeles, Los Angeles, California, United States of America.
Organizational Affiliation: