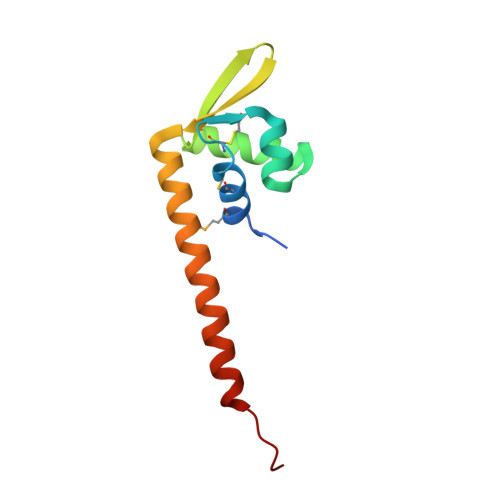
Diverse architectural properties of Sso10a proteins: Evidence for a role in chromatin compaction and organization.
Driessen, R.P., Lin, S.N., Waterreus, W.J., van der Meulen, A.L., van der Valk, R.A., Laurens, N., Moolenaar, G.F., Pannu, N.S., Wuite, G.J., Goosen, N., Dame, R.T.(2016) Sci Rep 6: 29422-29422
- PubMed: 27403582 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep29422
- Primary Citation Related Structures:
4HW0 - PubMed Abstract:
Sso10a proteins are small DNA-binding proteins expressed by the crenarchaeal model organism Sulfolobus solfataricus. Based on the structure of Sso10a1, which contains a winged helix-turn-helix motif, it is believed that Sso10a proteins function as sequence-specific transcription factors. Here we show that Sso10a1 and Sso10a2 exhibit different distinct DNA-binding modes. While the ability to bend DNA is shared between the two proteins, DNA bridging is observed only for Sso10a1 and only Sso10a2 exhibits filament formation along DNA. The architectural properties of Sso10a proteins suggest that these proteins fulfil generic roles in chromatin organization and compaction. As these proteins exhibit different binding behaviour depending on their DNA binding stoichiometry, altered levels of expression in the cell can be exploited to drive changes in local genome folding, which may operate to modulate transcription.
- Leiden Institute of Chemistry, Cell Observatory and Centre for Microbial Cell Biology, Leiden University, Einsteinweg 55, 2333 CC Leiden, The Netherlands.
Organizational Affiliation: