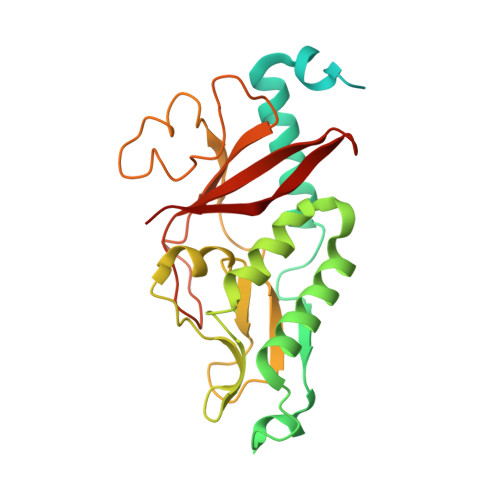
The type IV secretion protein TraK from the Enterococcus conjugative plasmid pIP501 exhibits a novel fold
Goessweiner-Mohr, N., Fercher, C., Arends, K., Birner-Gruenberger, R., Laverde-Gomez, D., Huebner, J., Grohmann, E., Keller, W.(2014) Acta Crystallogr D Biol Crystallogr 70: 1124-1135
- PubMed: 24699656 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004714001606
- Primary Citation Related Structures:
4HIC - PubMed Abstract:
Conjugative plasmid transfer presents a serious threat to human health as the most important means of spreading antibiotic resistance and virulence genes among bacteria. The required direct cell-cell contact is established by a multi-protein complex, the conjugative type IV secretion system (T4SS). The conjugative core complex spans the cellular envelope and serves as a channel for macromolecular secretion. T4SSs of Gram-negative (G-) origin have been studied in great detail. In contrast, T4SSs of Gram-positive (G+) bacteria have only received little attention thus far, despite the medical relevance of numerous G+ pathogens (e.g. enterococci, staphylococci and streptococci). This study provides structural information on the type IV secretion (T4S) protein TraK of the G+ broad host range Enterococcus conjugative plasmid pIP501. The crystal structure of the N-terminally truncated construct TraKΔ was determined to 3.0 Å resolution and exhibits a novel fold. Immunolocalization demonstrated that the protein localizes to the cell wall facing towards the cell exterior, but does not exhibit surface accessibility. Circular dichroism, dynamic light scattering and size-exclusion chromatography confirmed the protein to be a monomer. With the exception of proteins from closely related T4SSs, no significant sequence or structural relatives were found. This observation marks the protein as a very exclusive, specialized member of the pIP501 T4SS.
- Institute for Molecular Biosciences, Karl-Franzens-University Graz, Humboldtstrasse 50/III, 8010 Graz, Austria.
Organizational Affiliation: