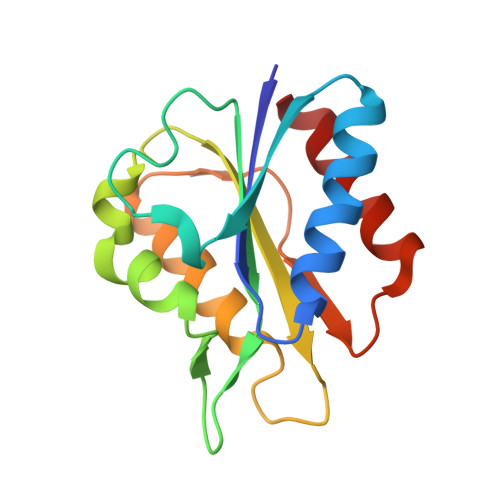
Crystal Structure of Dimeric Flavodoxin from Desulfovibrio gigas Suggests a Potential Binding Region for the Electron-Transferring Partner
Hsieh, Y.C., Chia, T.S., Fun, H.K., Chen, C.J.(2013) Int J Mol Sci 14: 1667-1683
- PubMed: 23322018 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms14011667
- Primary Citation Related Structures:
4HEQ - PubMed Abstract:
Flavodoxins, which exist widely in microorganisms, have been found in various pathways with multiple physiological functions. The flavodoxin (Fld) containing the cofactor flavin mononucleotide (FMN) from sulfur-reducing bacteria Desulfovibrio gigas (D. gigas) is a short-chain enzyme that comprises 146 residues with a molecular mass of 15 kDa and plays important roles in the electron-transfer chain. To investigate its structure, we purified this Fld directly from anaerobically grown D. gigas cells. The crystal structure of Fld, determined at resolution 1.3 Å, is a dimer with two FMN packing in an orientation head to head at a distance of 17 Å, which generates a long and connected negatively charged region. Two loops, Thr59-Asp63 and Asp95-Tyr100, are located in the negatively charged region and between two FMN, and are structurally dynamic. An analysis of each monomer shows that the structure of Fld is in a semiquinone state; the positions of FMN and the surrounding residues in the active site deviate. The crystal structure of Fld from D. gigas agrees with a dimeric form in the solution state. The dimerization area, dynamic characteristics and structure variations between monomers enable us to identify a possible binding area for its functional partners.
- Life Science Group, Scientific Research Division, National Synchrotron Radiation Research Center, Hsinchu 30076, Taiwan. cjchen@nsrrc.org.tw.
Organizational Affiliation: