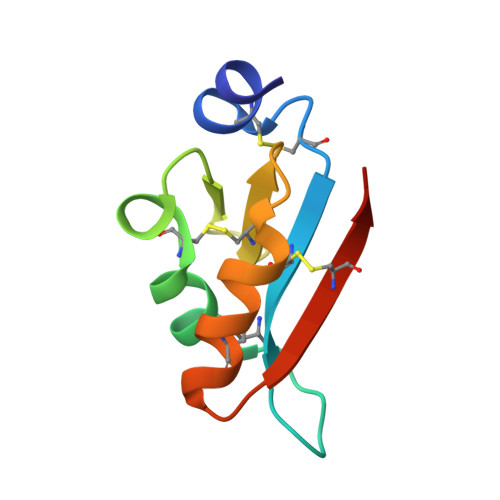
The Atomic Structure of the Virally Encoded Antifungal Protein, KP6.
Allen, A., Chatt, E., Smith, T.J.(2013) J Mol Biology 425: 609-621
- PubMed: 23219466 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2012.11.033
- Primary Citation Related Structures:
4GVB - PubMed Abstract:
Killer toxins are produced by several genera of yeast and filamentous fungi. A small proportion of Ustilago maydis strains produce killer toxins, to which they are resistant, but sensitive strains are the majority in the wild populations. There are three killer types (P1, P4 and P6) that secrete KP1, KP4 and KP6 toxins, respectively, which are produced only by strains persistently infected with double-stranded RNA viruses (UmV) in the cell cytoplasm. Unlike nearly all other viruses, UmV are only transmitted through mitosis or meiosis. As shown here, KP6 is different from any other known cytotoxic protein. KP6 is neutral protein composed of two subunits: KP6α and KP6β. KP6α is responsible for targeting while KP6β is cytotoxic. Neither subunit is homologous in either sequence or structure to any other toxin, but they have highly similar structures to each other. The major difference between the two subunits is a hydrophobic helix at the N-terminus of KP6α and is likely key to target recognition. Unlike any other toxin, KP6 is translated as a single polypeptide with a 31-residue linker region in the middle of the protein. From structural prediction studies, this linker likely makes for a more compact KP6 structure that sequesters the hydrophobic helix of KP6α. A model whereby the protoxin undergoes a conformational activation process that exposes this helix immediately prior to secretion is presented.
- Donald Danforth Plant Science Center, 975 North Warson Road, St. Louis, MO 63132, USA.
Organizational Affiliation: