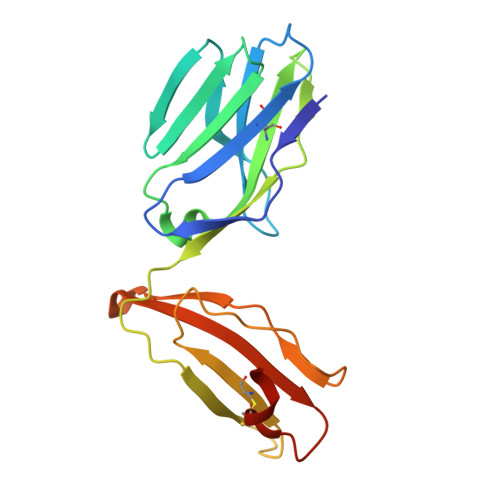
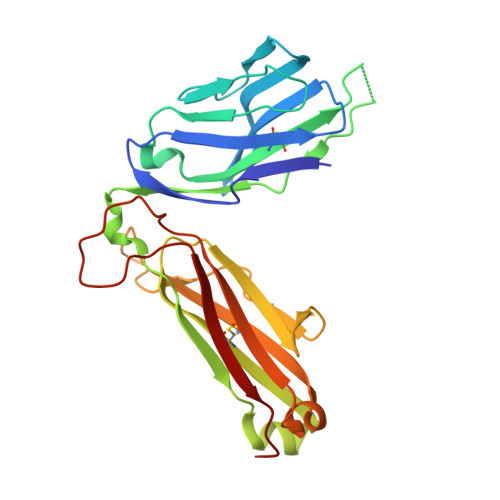
Increased Peptide Contacts Govern High Affinity Binding of a Modified TCR Whilst Maintaining a Native pMHC Docking Mode.
Cole, D.K., Sami, M., Scott, D.R., Rizkallah, P.J., Borbulevych, O.Y., Todorov, P.T., Moysey, R.K., Jakobsen, B.K., Boulter, J.M., Baker, B.M., Li, Y.I.(2013) Front Immunol 4: 168-168
- PubMed: 23805144 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fimmu.2013.00168
- Primary Citation Related Structures:
4GRM - PubMed Abstract:
Natural T cell receptors (TCRs) generally bind to their cognate pMHC molecules with weak affinity and fast kinetics, limiting their use as therapeutic agents. Using phage display, we have engineered a high affinity version of the A6 wild-type TCR (A6wt), specific for the human leukocyte antigen (HLA-A(∗)0201) complexed with human T cell lymphotropic virus type 111-19 peptide (A2-Tax). Mutations in just 4 residues in the CDR3β loop region of the A6wt TCR were selected that improved binding to A2-Tax by nearly 1000-fold. Biophysical measurements of this mutant TCR (A6c134) demonstrated that the enhanced binding was derived through favorable enthalpy and a slower off-rate. The structure of the free A6c134 TCR and the A6c134/A2-Tax complex revealed a native binding mode, similar to the A6wt/A2-Tax complex. However, concordant with the more favorable binding enthalpy, the A6c134 TCR made increased contacts with the Tax peptide compared with the A6wt/A2-Tax complex, demonstrating a peptide-focused mechanism for the enhanced affinity that directly involved the mutated residues in the A6c134 TCR CDR3β loop. This peptide-focused enhanced TCR binding may represent an important approach for developing antigen specific high affinity TCR reagents for use in T cell based therapies.
- Cardiff University School of Medicine, Heath Park , Cardiff , UK.
Organizational Affiliation: