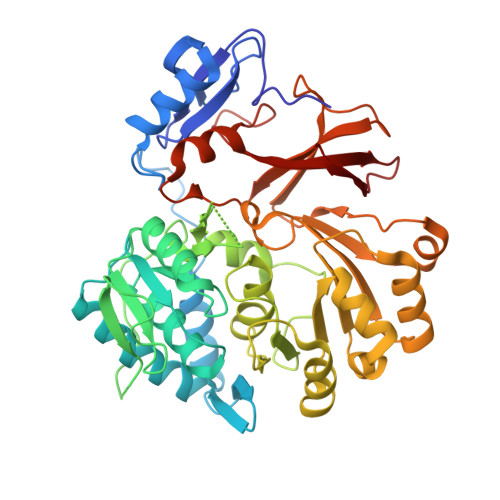
Structural Basis of the Interaction of MbtH-like Proteins, Putative Regulators of Nonribosomal Peptide Biosynthesis, with Adenylating Enzymes.
Herbst, D.A., Boll, B., Zocher, G., Stehle, T., Heide, L.(2013) J Biological Chem 288: 1991-2003
- PubMed: 23192349 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.420182
- Primary Citation Related Structures:
4GR4, 4GR5 - PubMed Abstract:
The biosynthesis of nonribosomally formed peptides (NRPs), which include important antibiotics such as vancomycin, requires the activation of amino acids through adenylate formation. The biosynthetic gene clusters of NRPs frequently contain genes for small, so-called MbtH-like proteins. Recently, it was discovered that these MbtH-like proteins are required for some of the adenylation reactions in NRP biosynthesis, but the mechanism of their interaction with the adenylating enzymes has remained unknown. In this study, we determined the structure of SlgN1, a 3-methylaspartate-adenylating enzyme involved in the biosynthesis of the hybrid polyketide/NRP antibiotic streptolydigin. SlgN1 contains an MbtH-like domain at its N terminus, and our analysis defines the parameters required for an interaction between MbtH-like domains and an adenylating enzyme. Highly conserved tryptophan residues of the MbtH-like domain critically contribute to this interaction. Trp-25 and Trp-35 form a cleft on the surface of the MbtH-like domain, which accommodates the alanine side chain of Ala-433 of the adenylating domain. Mutation of Ala-433 to glutamate abolished the activity of SlgN1. Mutation of Ser-23 of the MbtH-like domain to tyrosine resulted in strongly reduced activity. However, the activity of this S23Y mutant could be completely restored by addition of the intact MbtH-like protein CloY from another organism. This suggests that the interface found in the structure of SlgN1 is the genuine interface between MbtH-like proteins and adenylating enzymes.
- Interfakultäres Institut für Biochemie, Eberhard Karls-Universität Tübingen, 72076 Tübingen, Germany.
Organizational Affiliation: