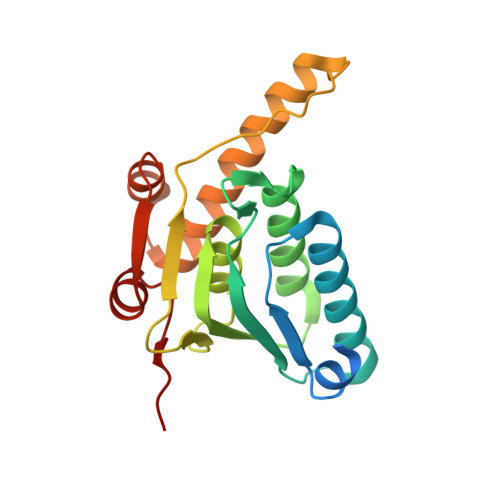
Structural Insights into the Inactive Subunit of the Apicoplast-localized Caseinolytic Protease Complex of Plasmodium falciparum.
El Bakkouri, M., Rathore, S., Calmettes, C., Wernimont, A.K., Liu, K., Sinha, D., Asad, M., Jung, P., Hui, R., Mohmmed, A., Houry, W.A.(2013) J Biological Chem 288: 1022-1031
- PubMed: 23192353 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.416560
- Primary Citation Related Structures:
4GM2, 4HNK - PubMed Abstract:
The ATP-dependent caseinolytic protease, ClpP, is highly conserved in bacteria and in the organelles of different organisms. In cyanobacteria, plant plastids, and the apicoplast of the genus Plasmodium, a noncatalytic paralog of ClpP, termed ClpR, has been identified. ClpRs are found to form heterocomplexes with ClpP resulting in a ClpRP tetradecameric cylinder having less than 14 catalytic triads. The exact role of ClpR in such a complex remains enigmatic. Here we describe the x-ray crystal structure of ClpR protein heptamer from Plasmodium falciparum (PfClpR). This is the first structure of a ClpR protein. The structure shows that the PfClpR monomer adopts a fold similar to that of ClpP, but has a unique motif, which we named the R-motif, forming a β turn located near the inactive catalytic triad in a three-dimensional space. The PfClpR heptamer exhibits a more open and flat ring than a ClpP heptamer. PfClpR was localized in the P. falciparum apicoplast as is the case of PfClpP. However, biochemical and structural data suggest that, contrary to what has been observed in other organisms, PfClpP and PfClpR do not form a stable heterocomplex in the apicoplast of P. falciparum.
- Department of Biochemistry, University of Toronto, Toronto, Ontario M5S 1A8, Canada.
Organizational Affiliation: