Optimization of Inhibitors of the Tyrosine Kinase EphB4. 2. Cellular Potency Improvement and Binding Mode Validation by X-ray Crystallography.
Lafleur, K., Dong, J., Huang, D., Caflisch, A., Nevado, C.(2013) J Med Chem 56: 84-96
- PubMed: 23253074 Search on PubMed
- DOI: https://doi.org/10.1021/jm301187e
- Primary Citation Related Structures:
4GK2, 4GK3, 4GK4 - PubMed Abstract:
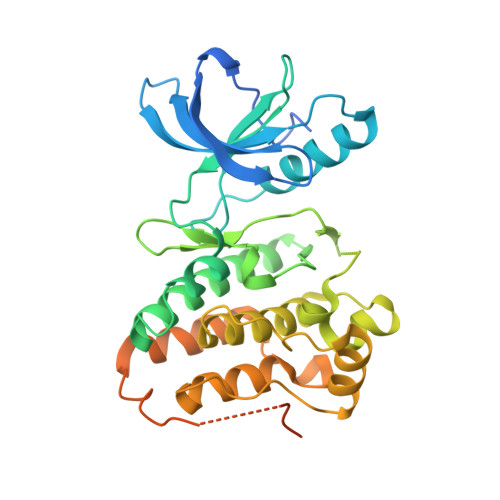
Inhibition of the tyrosine kinase erythropoietin-producing human hepatocellular carcinoma receptor B4 (EphB4) is an effective strategy for the treatment of solid tumors. We have previously reported a low nanomolar ATP-competitive inhibitor of EphB4 discovered in silico by fragment-based high-throughput docking combined with explicit solvent molecular dynamics simulations. Here we present a second generation of EphB4 inhibitors that show high inhibitory potency in both enzymatic and cell-based assays while preserving the appealing selectivity profile exhibited by the parent compound. In addition, respectable levels of antiproliferative activity for these compounds have been obtained. Finally, the binding mode predicted by docking and molecular dynamics simulations is validated by solving the crystal structures of three members of this chemical class in complex with the EphA3 tyrosine kinase whose ATP-binding site is essentially identical to that of EphB4.
- Department of Organic Chemistry, University of Zurich, Winterthurerstrasse 190, CH-8057 Zurich, Switzerland.
Organizational Affiliation: