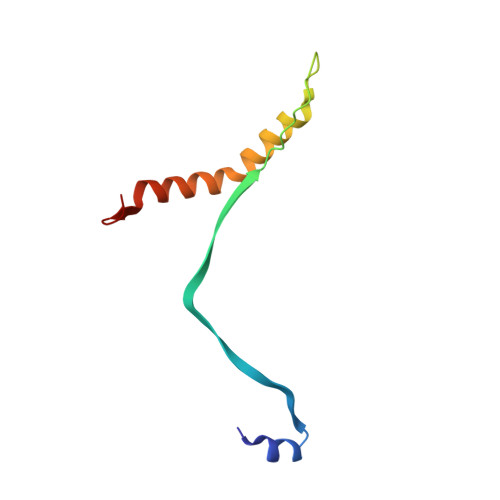
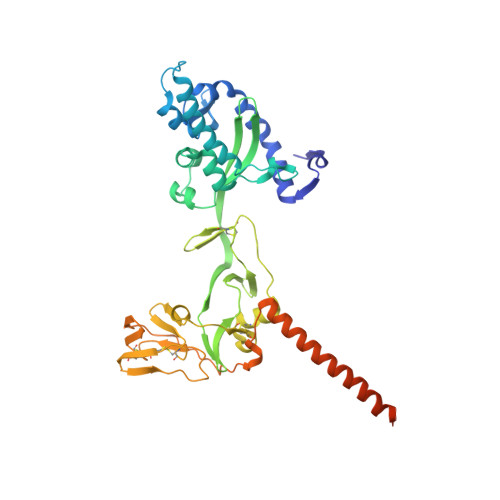
Structure of the cleavage-activated prefusion form of the parainfluenza virus 5 fusion protein.
Welch, B.D., Liu, Y., Kors, C.A., Leser, G.P., Jardetzky, T.S., Lamb, R.A.(2012) Proc Natl Acad Sci U S A 109: 16672-16677
- PubMed: 23012473 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1213802109
- Primary Citation Related Structures:
4GIP - PubMed Abstract:
The paramyxovirus parainfluenza virus 5 (PIV5) enters cells by fusion of the viral envelope with the plasma membrane through the concerted action of the fusion (F) protein and the receptor binding protein hemagglutinin-neuraminidase. The F protein folds initially to form a trimeric metastable prefusion form that is triggered to undergo large-scale irreversible conformational changes to form the trimeric postfusion conformation. It is thought that F refolding couples the energy released with membrane fusion. The F protein is synthesized as a precursor (F0) that must be cleaved by a host protease to form a biologically active molecule, F1,F2. Cleavage of F protein is a prerequisite for fusion and virus infectivity. Cleavage creates a new N terminus on F1 that contains a hydrophobic region, known as the FP, which intercalates target membranes during F protein refolding. The crystal structure of the soluble ectodomain of the uncleaved form of PIV5 F is known; here we report the crystal structure of the cleavage-activated prefusion form of PIV5 F. The structure shows minimal movement of the residues adjacent to the protease cleavage site. Most of the hydrophobic FP residues are buried in the uncleaved F protein, and only F103 at the newly created N terminus becomes more solvent-accessible after cleavage. The conformational freedom of the charged arginine residues that compose the protease recognition site increases on cleavage of F protein.
- Howard Hughes Medical Institute and Department of Molecular Biosciences, Northwestern University, Evanston, IL 60208, USA.
Organizational Affiliation: