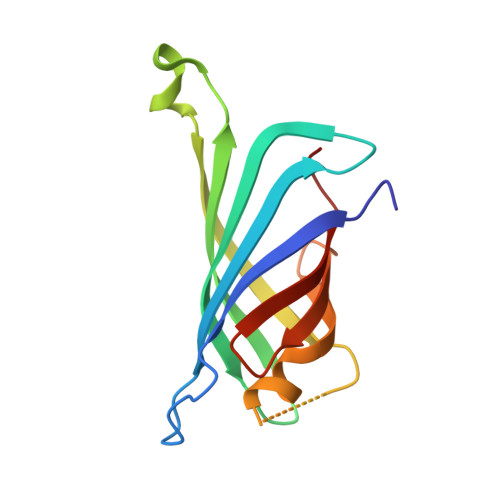
Structural consequences of cutting a binding loop: two circularly permuted variants of streptavidin.
Le Trong, I., Chu, V., Xing, Y., Lybrand, T.P., Stayton, P.S., Stenkamp, R.E.(2013) Acta Crystallogr D Biol Crystallogr 69: 968-977
- PubMed: 23695241 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444913003855
- Primary Citation Related Structures:
4GD9, 4GDA - PubMed Abstract:
Circular permutation of streptavidin was carried out in order to investigate the role of a main-chain amide in stabilizing the high-affinity complex of the protein and biotin. Mutant proteins CP49/48 and CP50/49 were constructed to place new N-termini at residues 49 and 50 in a flexible loop involved in stabilizing the biotin complex. Crystal structures of the two mutants show that half of each loop closes over the binding site, as observed in wild-type streptavidin, while the other half adopts the open conformation found in the unliganded state. The structures are consistent with kinetic and thermodynamic data and indicate that the loop plays a role in enthalpic stabilization of the bound state via the Asn49 amide-biotin hydrogen bond. In wild-type streptavidin, the entropic penalties of immobilizing a flexible portion of the protein to enhance binding are kept to a manageable level by using a contiguous loop of medium length (six residues) which is already constrained by its anchorage to strands of the β-barrel protein. A molecular-dynamics simulation for CP50/49 shows that cleavage of the binding loop results in increased structural fluctuations for Ser45 and that these fluctuations destabilize the streptavidin-biotin complex.
- Department of Biological Structure, University of Washington, Box 357420, Seattle, WA 98195-7420, USA.
Organizational Affiliation: