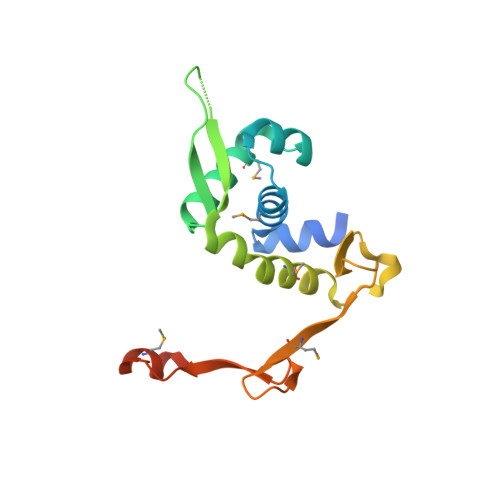
The X-ray crystal structure of PA1374 from Pseudomonas aeruginosa, a putative oxidative-stress sensing transcriptional regulator.
Kim, H., Choe, J.(2013) Biochem Biophys Res Commun 431: 376-381
- PubMed: 23337505 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2013.01.044
- Primary Citation Related Structures:
4GCV - PubMed Abstract:
Members of the multiple antibiotic resistance regulator (MarR) family regulate the expression of genes related to antibiotic resistance, oxidative stress, and virulence in bacteria and Archaea. Here, we determined the structure of PA1374 from Pseudomonas aeruginosa at 2.3Å resolution. PA1374 belonged to the MarR family and its structure revealed a tightly bound dimer with each subunit containing a winged helix-turn-helix (wHTH) DNA-binding motif. Conserved arginine residues, Arg55, Arg74, and Arg77, were located in the wHTH region, which might be important to DNA binding. Furthermore, each monomer contained a pocket made of conserved hydrophobic residues. A highly conserved Cys11 located at one end of this pocket may undergo oxidation by organic hydroperoxide molecules, as shown in other MarR family proteins acting as redox-sensing regulators. These results provide insights about the role of PA1374 as a putative oxidative-stress sensing transcriptional regulator.
- Department of Life Science, University of Seoul, Seoul 130-743, Republic of Korea.
Organizational Affiliation: