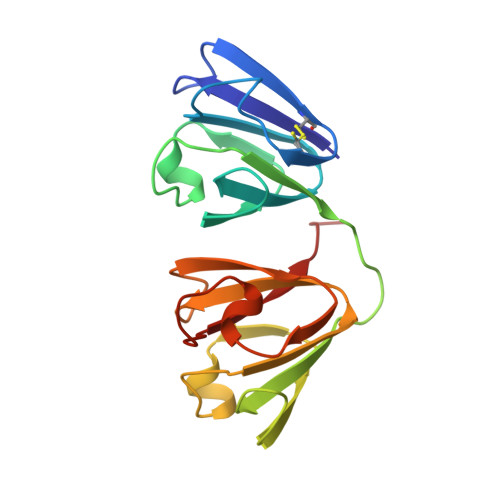
Structure of the bovine eye lens protein gammaB(gammaII)-crystallin at 1.47 A.
Najmudin, S., Nalini, V., Driessen, H.P., Slingsby, C., Blundell, T.L., Moss, D.S., Lindley, P.F.(1993) Acta Crystallogr D Biol Crystallogr 49: 223-233
- PubMed: 15299528
- DOI: https://doi.org/10.1107/S0907444992007601
- Primary Citation of Related Structures:
4GCR - PubMed Abstract:
The molecular structure of calf gammaB-crystallin (previously called gammaII), a lens-specific protein, has been refined to a crystallographic R factor of 18.1% for all reflection data, between 8.0 and 1.47 A, 25 959 hkl measured at 293 (1) K. 230 water molecules have been defined by difference Fourier techniques and included in a restrained least-squares refinement. Difference Fourier maps clearly indicated the presence of multiple sites for the sulfur atoms of Cys 18 and Cys 22 which were therefore given coupled second-site occupancies during the refinement. The sulfur atom in the major position of Cys 22 is in the reduced state. Either of the Cys 18 sites can form a high-energy disulfide bridge with the minor position of Cys 22. The position of the carboxy terminus and many other surface side chains have been further defined including the RGD signal peptide. The hydration of the backbone and the interdomain region has been analysed. 27 water molecules make extensive contacts to a single protein molecule and thus contribute to its stability.
- Laboratory of Molecular Biology, Department of Crystallography, Birkbeck College, London, England.
Organizational Affiliation: