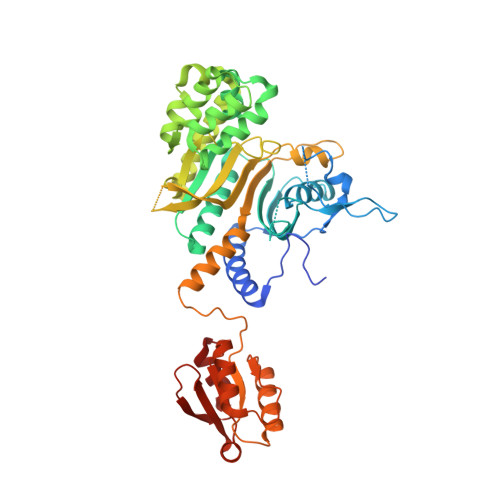
Internally Deleted Human tRNA Synthetase Suggests Evolutionary Pressure for Repurposing.
Xu, Z., Wei, Z., Zhou, J.J., Ye, F., Lo, W.S., Wang, F., Lau, C.F., Wu, J., Nangle, L.A., Chiang, K.P., Yang, X.L., Zhang, M., Schimmel, P.(2012) Structure 20: 1470-1477
- PubMed: 22958643 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2012.08.001
- Primary Citation Related Structures:
4G84, 4G85 - PubMed Abstract:
Aminoacyl-tRNA synthetases (AARSs) catalyze aminoacylation of tRNAs in the cytoplasm. Surprisingly, AARSs also have critical extracellular and nuclear functions. Evolutionary pressure for new functions might be manifested by splice variants that skip only an internal catalytic domain (CD) and link noncatalytic N- and C-terminal polypeptides. Using disease-associated histidyl-tRNA synthetase (HisRS) as an example, we found an expressed 171-amino acid protein (HisRSΔCD) that deleted the entire CD, and joined an N-terminal WHEP to the C-terminal anticodon-binding domain (ABD). X-ray crystallography and three-dimensional NMR revealed the structures of human HisRS and HisRSΔCD. In contrast to homodimeric HisRS, HisRSΔCD is monomeric, where rupture of the ABD's packing with CD resulted in a dumbbell-like structure of flexibly linked WHEP and ABD domains. In addition, the ABD of HisRSΔCD presents a distinct local conformation. This natural internally deleted HisRS suggests evolutionary pressure to reshape AARS tertiary and quaternary structures for repurposing.
- IAS HKUST-Scripps R&D Laboratory, Institute for Advanced Study, Hong Kong University of Science and Technology, Clear Water Bay, Kowloon, Hong Kong, China.
Organizational Affiliation: