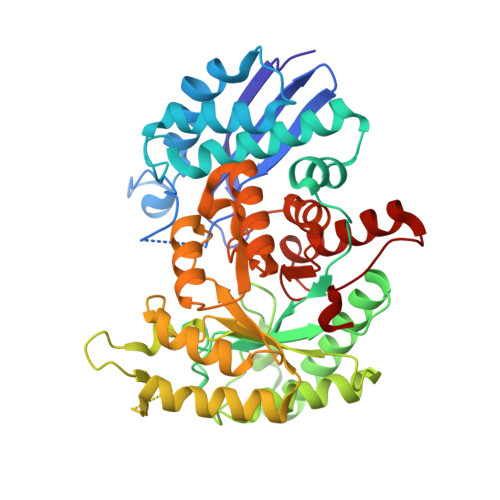
Structural Characterization of Glycolytic Enzymes from Trypanosoma cruzi.
Austin, K., Obachi, V.A., Muzenda, F.L., Moetlediwa, M.T., Agyei, C., Craig, T., Abendroth, J., Edwards, T., Nguyen, M., Tran, N., Staker, B., Subramanian, S., Myler, P., Zininga, T., Govender, K.K., Chakafana, G.(2026) Mol Biochem Parasitol : 111736-111736
- PubMed: 41713750 Search on PubMed
- DOI: https://doi.org/10.1016/j.molbiopara.2026.111736
- Primary Citation Related Structures:
4G7F, 4QFH - PubMed Abstract:
Trypanosoma cruzi, the etiological agent of Chagas disease, depends on glycolysis for ATP production, rendering its glycolytic enzymes attractive targets for therapeutic development. Here, we report the high-resolution crystal structures of two essential glycolytic enzymes, glucose-6-phosphate isomerase (Tc PGI, 1.8 Å) and enolase (Tc enolase, 2.4 Å) and provide structural and computational analyses to support structure-based drug design. Tc PGI adopts a dimeric αβα sandwich fold and features a parasite-specific 53-residue N-terminal extension and a unique C-terminal hook region which both distinguish it from its human ortholog. Tc enolase exhibits the conserved (α/β) 8 TIM barrel fold but harbors minor distinct structural deviations, including an extended α17 helix and a structured α1 region, which differentiate it from human isoforms. Both enzymes exhibited high thermal stability, consistent with adaptation to the parasite's complex life cycle. Structure-based virtual screening using a scaffold with known multi-target potential identified distinct high-affinity inhibitors for each enzyme. Molecular dynamics simulations further confirmed stable enzyme-inhibitor interactions and favorable binding energetics. Collectively, these findings reveal structural signatures unique to T. cruzi glycolytic enzymes and lay the groundwork for the development of antiparasitic therapeutics.
- Chemistry and Biochemistry Department, Hampton University, 200 William R Harvey Way, Hampton, VA 23668, USA.
Organizational Affiliation: