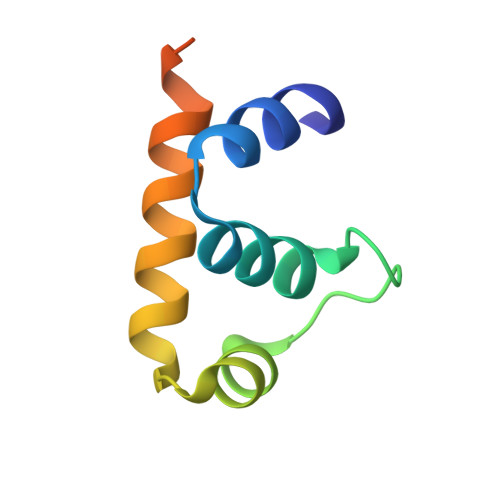
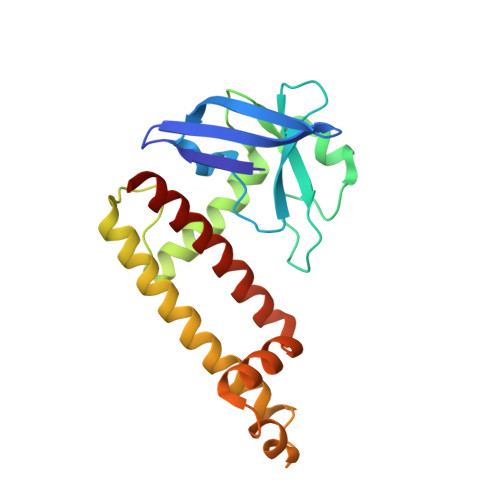
Promoter-specific transcription inhibition in Staphylococcus aureus by a phage protein.
Osmundson, J., Montero-Diez, C., Westblade, L.F., Hochschild, A., Darst, S.A.(2012) Cell 151: 1005-1016
- PubMed: 23178120
- DOI: https://doi.org/10.1016/j.cell.2012.10.034
- Primary Citation of Related Structures:
4G6D, 4G8X, 4G94 - PubMed Abstract:
Phage G1 gp67 is a 23 kDa protein that binds to the Staphylococcus aureus (Sau) RNA polymerase (RNAP) σ(A) subunit and blocks cell growth by inhibiting transcription. We show that gp67 has little to no effect on transcription from most promoters but is a potent inhibitor of ribosomal RNA transcription. A 2.0-Å-resolution crystal structure of the complex between gp67 and Sau σ(A) domain 4 (σ(A)(4)) explains how gp67 joins the RNAP promoter complex through σ(A)(4) without significantly affecting σ(A)(4) function. Our results indicate that gp67 forms a complex with RNAP at most, if not all, σ(A)-dependent promoters, but selectively inhibits promoters that depend on an interaction between upstream DNA and the RNAP α-subunit C-terminal domain (αCTD). Thus, we reveal a promoter-specific transcription inhibition mechanism by which gp67 interacts with the RNAP promoter complex through one subunit (σ(A)), and selectively affects the function of another subunit (αCTD) depending on promoter usage.
- The Rockefeller University, 1230 York Avenue, New York, NY 10065, USA.
Organizational Affiliation: