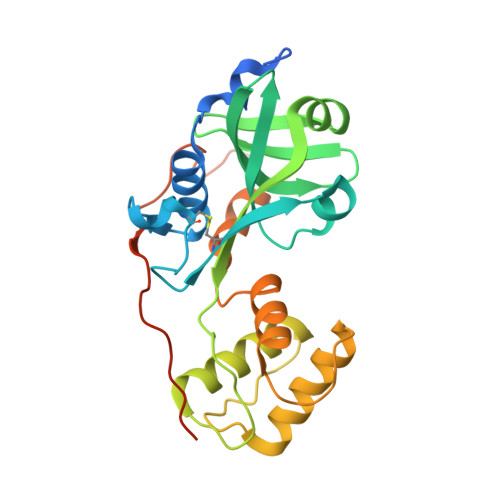
Structure of a periplasmic domain of the EpsAB fusion protein of the Vibrio vulnificus type II secretion system.
Martynowski, D., Grochulski, P., Howard, P.S.(2013) Acta Crystallogr D Biol Crystallogr 69: 142-149
- PubMed: 23385451
- DOI: https://doi.org/10.1107/S0907444912042710
- Primary Citation Related Structures:
4G54 - PubMed Abstract:
Vibrio vulnificus utilizes the type II secretion system (T2SS), culminating in a megadalton outer membrane complex called the secretin, to translocate extracellular proteins from the periplasmic space across the outer membrane. In Aeromonas hydrophila, the general secretion pathway proteins ExeA and ExeB form an inner membrane complex which interacts with peptidoglycan and is required for the assembly of the secretin composed of ExeD. In V. vulnificus, these two proteins are fused into one protein, EpsAB. Here, the crystal structure of a periplasmic domain of EpsAB (amino acids 333-584) solved by SAD phasing is presented. The crystals belonged to space group C2 and diffracted to 1.55 Å resolution.
- Department of Microbiology and Immunology, University of Saskatchewan, 107 Wiggins Road, Saskatoon, SK S7N 2W8, Canada.
Organizational Affiliation: